TRA2β 的自动调节剪接可在抗原受体刺激下对 T 细胞的命运进行调控
IF 44.7
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
摘要
T细胞受体(TCR)对多肽-主要组织相容性复合体(MHC)的敏感性决定了T细胞的命运。TCR敏感性的经典模型不能完全用转录调控来解释。在这项研究中,我们发现了一种 TCR 敏感性的转录后调控机制,该机制通过小鼠和人类 RNA 结合蛋白(RBP)TRA2β 中一个进化超保守的毒外显子(PE)引导 TCR 信号转录本的替代剪接。在癌症和感染期间出现的 TRA2β-PE 剪接是 TCR 诱导的效应 T 细胞扩增和功能所必需的。跳过TRA2β-PE可提高TCR的敏感性,从而增强T细胞对抗原的反应。随着抗原水平的降低,Tra2β-PE 的重新整合使 T 细胞得以存活。最后,我们发现 TRA2β-PE 首次出现在有颌脊椎动物的基因组中,这些脊椎动物能够进行 TCR 基因重排。我们认为,TRA2β-PE的剪接是TCR敏感性的看门人,决定着T细胞的命运。本文章由计算机程序翻译,如有差异,请以英文原文为准。
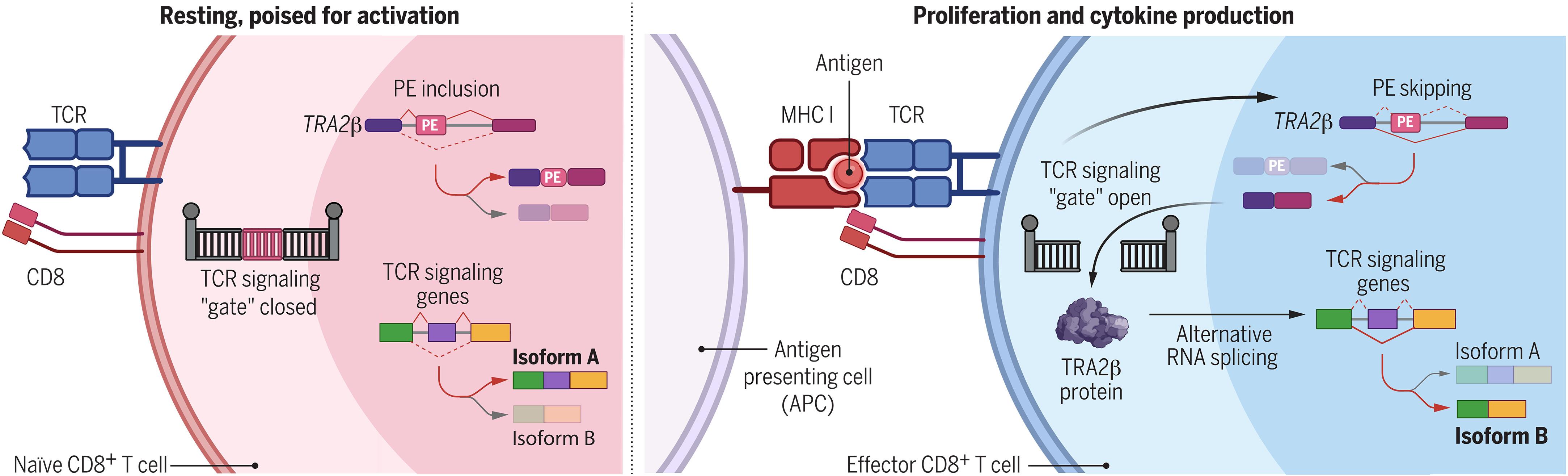
Autoregulated splicing of TRA2β programs T cell fate in response to antigen-receptor stimulation
T cell receptor (TCR) sensitivity to peptide–major histocompatibility complex (MHC) dictates T cell fate. Canonical models of TCR sensitivity cannot be fully explained by transcriptional regulation. In this work, we identify a posttranscriptional regulatory mechanism of TCR sensitivity that guides alternative splicing of TCR signaling transcripts through an evolutionarily ultraconserved poison exon (PE) in the RNA-binding protein (RBP) TRA2β in mouse and human. TRA2β-PE splicing, seen during cancer and infection, was required for TCR-induced effector T cell expansion and function. Tra2β-PE skipping enhanced T cell response to antigen by increasing TCR sensitivity. As antigen levels decreased, Tra2β-PE reinclusion allowed T cell survival. Finally, we found that TRA2β-PE was first included in the genome of jawed vertebrates that were capable of TCR gene rearrangements. We propose that TRA2β-PE splicing acts as a gatekeeper of TCR sensitivity to shape T cell fate.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Science
综合性期刊-综合性期刊
CiteScore
61.10
自引率
0.90%
发文量
0
审稿时长
2.1 months
期刊介绍:
Science is a leading outlet for scientific news, commentary, and cutting-edge research. Through its print and online incarnations, Science reaches an estimated worldwide readership of more than one million. Science’s authorship is global too, and its articles consistently rank among the world's most cited research.
Science serves as a forum for discussion of important issues related to the advancement of science by publishing material on which a consensus has been reached as well as including the presentation of minority or conflicting points of view. Accordingly, all articles published in Science—including editorials, news and comment, and book reviews—are signed and reflect the individual views of the authors and not official points of view adopted by AAAS or the institutions with which the authors are affiliated.
Science seeks to publish those papers that are most influential in their fields or across fields and that will significantly advance scientific understanding. Selected papers should present novel and broadly important data, syntheses, or concepts. They should merit recognition by the wider scientific community and general public provided by publication in Science, beyond that provided by specialty journals. Science welcomes submissions from all fields of science and from any source. The editors are committed to the prompt evaluation and publication of submitted papers while upholding high standards that support reproducibility of published research. Science is published weekly; selected papers are published online ahead of print.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


