Development and Comprehensive Evaluation of Culture-Independent, Long Amplicon-Based Targeted Next-Generation Sequencing Methods for Predicting Antimicrobial Resistance in Tuberculosis
IF 6.7
1区 化学
Q1 CHEMISTRY, ANALYTICAL
引用次数: 0
Abstract
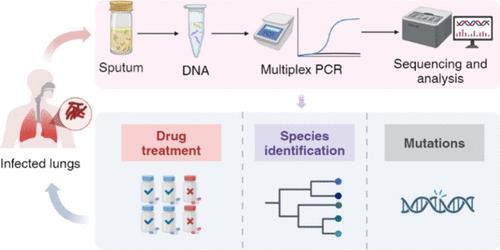
The great variety of antimicrobial resistance (AMR) profiles among tuberculosis (TB) patients necessitates a comprehensive detection method. This study developed culture-independent, long amplicon-based targeted next-generation sequencing (tNGS) methods for predicting AMR across 16 drugs within the Mycobacterium tuberculosis complex (MTBC). Multiplex PCR amplification was employed to enrich 20 gene regions, with sequencing performed on either the Oxford Nanopore Technologies (ONT) or Illumina platforms. Customized bioinformatics pipelines provide a streamlined process from raw data to clinician-friendly reports. The ONT tNGS method has been optimized, and its performance has been thoroughly evaluated, utilizing Q20+ chemistry in combination with the R10.4.1 flow cell. It requires only 15 high-quality reads per target gene to accurately identify all variants, with the turnaround time taking 4 h and 50 min. Studies confirmed that this method effectively identifies Mycobacterium species and was highly resistant to interference from other clinical pathogens. To ensure optimal coverage, it is recommended to input at least 500 copies of the genome and sequence 500MB of high-quality FASTQ data. Diagnostic performance evaluations demonstrate that this method achieves 98.35% concordance with phenotypic drug susceptibility testing (pDST) and is consistent with the results obtained from Xpert MTB/RIF assays. The design of long amplicons not only ensures comprehensive coverage of target regions but also simplifies primer design, facilitating compatibility with various sequencing platforms. Compared with previous studies, the optimized ONT tNGS method in this study significantly improves turnaround time, detection accuracy, and the comprehensive coverage of mutations associated with AMR.
开发和全面评估基于长扩增片段的培养依赖性靶向新一代测序方法,用于预测结核病的抗菌药耐药性
结核病患者抗菌素耐药性(AMR)谱的多样性需要一种综合的检测方法。该研究开发了不依赖培养的、基于长扩增子的靶向下一代测序(tNGS)方法,用于预测结核分枝杆菌复合体(MTBC)内16种药物的AMR。采用多重PCR扩增富集20个基因区域,测序在Oxford Nanopore Technologies (ONT)或Illumina平台上进行。定制的生物信息学管道提供了从原始数据到临床友好报告的简化过程。利用Q20+化学与R10.4.1液流池相结合,对ONT tNGS方法进行了优化,并对其性能进行了全面评估。每个目标基因只需要15个高质量的reads就能准确识别所有变异,周转时间为4小时50分钟。研究证实,该方法可以有效识别分枝杆菌种类,并且对其他临床病原体的干扰具有很强的抵抗力。为确保最佳覆盖,建议至少输入500个基因组拷贝,并对500MB高质量FASTQ数据进行测序。诊断性能评价表明,该方法与表型药敏试验(pDST)的一致性达到98.35%,与Xpert MTB/RIF试验结果一致。长扩增子的设计不仅保证了目标区域的全面覆盖,而且简化了引物设计,便于与各种测序平台兼容。与以往的研究相比,本研究优化的ONT tNGS方法显著提高了周转时间、检测精度和AMR相关突变的全面覆盖率。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Analytical Chemistry
化学-分析化学
CiteScore
12.10
自引率
12.20%
发文量
1949
审稿时长
1.4 months
期刊介绍:
Analytical Chemistry, a peer-reviewed research journal, focuses on disseminating new and original knowledge across all branches of analytical chemistry. Fundamental articles may explore general principles of chemical measurement science and need not directly address existing or potential analytical methodology. They can be entirely theoretical or report experimental results. Contributions may cover various phases of analytical operations, including sampling, bioanalysis, electrochemistry, mass spectrometry, microscale and nanoscale systems, environmental analysis, separations, spectroscopy, chemical reactions and selectivity, instrumentation, imaging, surface analysis, and data processing. Papers discussing known analytical methods should present a significant, original application of the method, a notable improvement, or results on an important analyte.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


