Raman Spectral Feature Enhancement Framework for Complex Multiclassification Tasks
IF 6.7
1区 化学
Q1 CHEMISTRY, ANALYTICAL
引用次数: 0
Abstract
Raman spectroscopy enables label-free clinical diagnosis in a single step. However, identifying an individual carrying a specific disease from people with a multi-disease background is challenging. To address this, we developed a Raman spectral implicit feature augmentation with a Raman Intersection, Union, and Subtraction augmentation strategy (RIUS). RIUS expands the data set without requiring additional labeled data by leveraging set operations at the feature level, significantly enhancing model performance across various applications. On a challenging 30-class bacterial classification task, RIUS demonstrated a substantial improvement, increasing the accuracy of ResNet by 2.1% and that of SE-ResNet by 1.4%, achieving accuracies of 85.7% and 87.1%, respectively, on the Bacteria-ID-4 Data set, where RIUS improved ResNet and SE-ResNet accuracies by 13.6% and 14.5%, respectively, with only ten samples per category. When the sample size was reduced, accuracy gains increased to 31.7% and 38.3%, demonstrating the method’s robustness across different sample volumes. Compared to basic augmentation, our method exhibited superior performance across various sample volumes and demonstrated exceptional adaptability to different levels of complexity. RIUS exhibited superior performance, particularly in complex settings. Moreover, cluster analysis validated the effectiveness of the implicit feature augmentation module and the consistency between theoretical design and experimental results. We further validated our approach using clinical serum samples from 70 breast cancer patients and 70 controls, achieving an AUC of 0.94 and a sensitivity of 92.9%. Our approach enhances the potential for precisely identifying diseases in complex settings and offers plug-and-play enhancement for existing classification models.
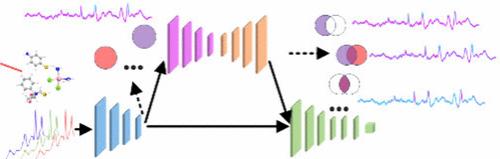
复杂多分类任务的拉曼光谱特征增强框架
拉曼光谱可以在一个步骤中进行无标签的临床诊断。然而,从具有多种疾病背景的人群中识别携带特定疾病的个体是具有挑战性的。为了解决这个问题,我们开发了一种拉曼光谱隐式特征增强方法,该方法采用拉曼交、并、减增强策略(RIUS)。RIUS扩展了数据集,而不需要额外的标记数据,通过利用特征级的集合操作,显著提高了各种应用程序的模型性能。在具有挑战性的30类细菌分类任务中,RIUS表现出了显著的改进,将ResNet的准确率提高了2.1%,将SE-ResNet的准确率提高了1.4%,在Bacteria-ID-4数据集上分别达到了85.7%和87.1%,其中RIUS将ResNet和SE-ResNet的准确率分别提高了13.6%和14.5%,每个类别只有10个样本。当样本量减少时,准确率提高到31.7%和38.3%,表明该方法在不同样本量下具有稳健性。与基本增强方法相比,我们的方法在不同的样品体积上表现出优异的性能,并表现出对不同复杂程度的卓越适应性。RIUS表现出优异的性能,特别是在复杂环境中。聚类分析验证了隐式特征增强模块的有效性以及理论设计与实验结果的一致性。我们使用来自70名乳腺癌患者和70名对照者的临床血清样本进一步验证了我们的方法,AUC为0.94,灵敏度为92.9%。我们的方法提高了在复杂环境中精确识别疾病的潜力,并为现有的分类模型提供了即插即用的增强。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Analytical Chemistry
化学-分析化学
CiteScore
12.10
自引率
12.20%
发文量
1949
审稿时长
1.4 months
期刊介绍:
Analytical Chemistry, a peer-reviewed research journal, focuses on disseminating new and original knowledge across all branches of analytical chemistry. Fundamental articles may explore general principles of chemical measurement science and need not directly address existing or potential analytical methodology. They can be entirely theoretical or report experimental results. Contributions may cover various phases of analytical operations, including sampling, bioanalysis, electrochemistry, mass spectrometry, microscale and nanoscale systems, environmental analysis, separations, spectroscopy, chemical reactions and selectivity, instrumentation, imaging, surface analysis, and data processing. Papers discussing known analytical methods should present a significant, original application of the method, a notable improvement, or results on an important analyte.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


