Detection of Inverse Stable Isotopic Labeling in Untargeted Metabolomic Data
IF 6.7
1区 化学
Q1 CHEMISTRY, ANALYTICAL
引用次数: 0
Abstract
Stable isotopic labeling is a powerful tool for determining the biosynthetic origin of metabolites and for discovering natural products that incorporate precursors of interest. When isotopically substituted precursors are not available commercially or synthetically, inverse stable isotopic labeling (InverSIL) is a useful alternative. With InverSIL, an organism is grown on an isotopically substituted medium and then fed precursors of natural isotopic abundance which can be tracked by mass spectrometry, thereby bypassing issues with precursor availability. Currently, there is no automated way to identify precursor incorporation in untargeted metabolomic data using InverSIL without specifying an expected change in the mass-to-charge ratio of metabolites that have incorporated the precursor. This makes it difficult to identify unknown natural products that may incorporate portions of precursors of interest using new biochemistry or to rapidly identify incorporation of multiple precursors into different metabolites simultaneously. To address this, we developed a new, robust workflow for the automated identification of inverse labeling in untargeted metabolomic data. We then use this method to identify metabolites that incorporate para-aminobenzoic acid and different portions of l-methionine, including in the same sample, and in the process discover the likely biosynthetic origin for the C-7 and C-9 methyl groups of the pterin portion of dephosphotetrahydromethanopterin, a C1 transfer coenzyme used by methylotrophic bacteria. This workflow can be applied in the future to streamline the use of the versatile InverSIL approach for natural product and metabolism research.
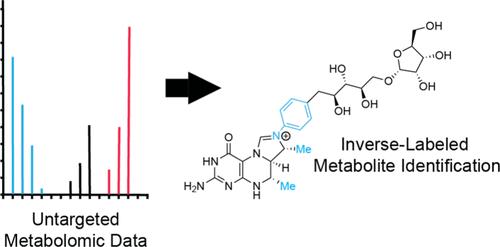
检测非目标代谢组数据中的反稳定同位素标记
稳定同位素标记是确定代谢物生物合成来源和发现含有相关前体的天然产物的有力工具。当无法从商业或合成途径获得同位素取代的前体时,反向稳定同位素标记(InverSIL)是一种有用的替代方法。使用反稳定同位素标记法,生物体在同位素替代的培养基上生长,然后向其输入天然同位素丰度的前体,这些前体可通过质谱法进行跟踪,从而避免了前体可用性的问题。目前,使用 InverSIL 还无法自动识别非靶向代谢组数据中的前体掺入情况,而无需指定掺入前体的代谢物质量电荷比的预期变化。这样就很难利用新的生物化学方法识别可能掺入了部分感兴趣前体的未知天然产物,也很难同时快速识别多种前体掺入不同代谢物的情况。为了解决这个问题,我们开发了一种新的、强大的工作流程,用于自动识别非靶向代谢组数据中的反向标记。然后,我们利用这种方法识别了在同一样本中结合对氨基苯甲酸和不同部分 l-蛋氨酸的代谢物,并在这一过程中发现了脱磷四氢甲蝶呤的 C-7 和 C-9 甲基的可能生物合成来源,脱磷四氢甲蝶呤是养甲细菌使用的一种 C1 转移辅酶。这一工作流程今后可用于简化天然产物和新陈代谢研究中多功能 InverSIL 方法的使用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Analytical Chemistry
化学-分析化学
CiteScore
12.10
自引率
12.20%
发文量
1949
审稿时长
1.4 months
期刊介绍:
Analytical Chemistry, a peer-reviewed research journal, focuses on disseminating new and original knowledge across all branches of analytical chemistry. Fundamental articles may explore general principles of chemical measurement science and need not directly address existing or potential analytical methodology. They can be entirely theoretical or report experimental results. Contributions may cover various phases of analytical operations, including sampling, bioanalysis, electrochemistry, mass spectrometry, microscale and nanoscale systems, environmental analysis, separations, spectroscopy, chemical reactions and selectivity, instrumentation, imaging, surface analysis, and data processing. Papers discussing known analytical methods should present a significant, original application of the method, a notable improvement, or results on an important analyte.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


