通过矩阵和张量分解无监督的基于EHR的表型
IF 11.7
2区 计算机科学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
Wiley Interdisciplinary Reviews-Data Mining and Knowledge Discovery
Pub Date : 2022-09-01
DOI:10.1002/widm.1494
引用次数: 2
摘要
计算表型允许从电子健康记录(EHR)中无监督地发现患者亚组以及相应的共同发生的医疗状况。通常,电子病历数据包含人口统计信息、诊断和实验室结果。发现(新的)表型具有潜在的预后和治疗价值。为医疗从业者提供透明、可解释的结果是推进精准医疗的重要要求和重要组成部分。低秩数据近似方法,如矩阵(例如,非负矩阵分解)和张量分解(例如,CANDECOMP/PARAFAC)已经证明它们可以提供这样透明和可解释的见解。最近的发展已经通过结合不同的约束和正则化来适应低秩数据近似方法,从而进一步促进可解释性。此外,它们还为EHR数据中的常见挑战(如高维、数据稀疏和不完整)提供了解决方案。特别是从纵向电子病历中提取时间表型近年来受到广泛关注。在本文中,我们对基于低秩近似的计算表型方法进行了全面的综述。现有文献分为基于矩阵与张量分解的时间与静态表型方法。此外,我们概述了不同的方法来验证表型,即临床意义的评估。本文章由计算机程序翻译,如有差异,请以英文原文为准。
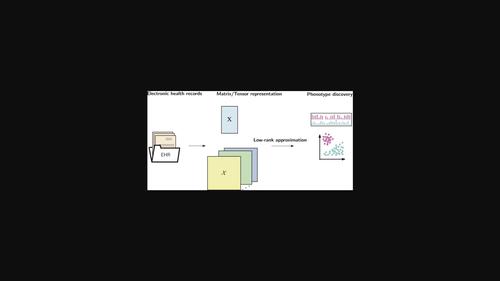
Unsupervised EHR‐based phenotyping via matrix and tensor decompositions
Computational phenotyping allows for unsupervised discovery of subgroups of patients as well as corresponding co‐occurring medical conditions from electronic health records (EHR). Typically, EHR data contains demographic information, diagnoses and laboratory results. Discovering (novel) phenotypes has the potential to be of prognostic and therapeutic value. Providing medical practitioners with transparent and interpretable results is an important requirement and an essential part for advancing precision medicine. Low‐rank data approximation methods such as matrix (e.g., nonnegative matrix factorization) and tensor decompositions (e.g., CANDECOMP/PARAFAC) have demonstrated that they can provide such transparent and interpretable insights. Recent developments have adapted low‐rank data approximation methods by incorporating different constraints and regularizations that facilitate interpretability further. In addition, they offer solutions for common challenges within EHR data such as high dimensionality, data sparsity and incompleteness. Especially extracting temporal phenotypes from longitudinal EHR has received much attention in recent years. In this paper, we provide a comprehensive review of low‐rank approximation‐based approaches for computational phenotyping. The existing literature is categorized into temporal versus static phenotyping approaches based on matrix versus tensor decompositions. Furthermore, we outline different approaches for the validation of phenotypes, that is, the assessment of clinical significance.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Wiley Interdisciplinary Reviews-Data Mining and Knowledge Discovery
COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE-COMPUTER SCIENCE, THEORY & METHODS
CiteScore
22.70
自引率
2.60%
发文量
39
审稿时长
>12 weeks
期刊介绍:
The goals of Wiley Interdisciplinary Reviews-Data Mining and Knowledge Discovery (WIREs DMKD) are multifaceted. Firstly, the journal aims to provide a comprehensive overview of the current state of data mining and knowledge discovery by featuring ongoing reviews authored by leading researchers. Secondly, it seeks to highlight the interdisciplinary nature of the field by presenting articles from diverse perspectives, covering various application areas such as technology, business, healthcare, education, government, society, and culture. Thirdly, WIREs DMKD endeavors to keep pace with the rapid advancements in data mining and knowledge discovery through regular content updates. Lastly, the journal strives to promote active engagement in the field by presenting its accomplishments and challenges in an accessible manner to a broad audience. The content of WIREs DMKD is intended to benefit upper-level undergraduate and postgraduate students, teaching and research professors in academic programs, as well as scientists and research managers in industry.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


