一种预测肝细胞癌患者预后和免疫治疗效果的杯状相关LncRNA风险模型。
摘要
铜中毒是一种新的程序性细胞死亡途径,由铜与脂酰化三羧酸(TCA)循环蛋白的直接结合引发。最近的研究表明,铜中毒相关基因调节肿瘤的发生。然而,铜中毒相关的长非编码RNA(lncRNA)在肝细胞癌(HCC)中的潜在作用和临床意义尚未确定。我们对从癌症基因组图谱(TCGA)数据集中提取的HCC患者的RNA序列数据进行了生物信息学分析,以确定和验证铜中毒相关lncRNA预后标志。此外,我们分析了铜中毒相关lncRNA的预后标志在预测免疫治疗效果和肿瘤免疫微环境状况方面的临床意义。从TCGA肝细胞癌(TCGA-LIHC)数据集中下载374个HCC样本和50个正常肝脏样本的RNA测序数据、基因组突变和临床信息。基因lncRNA对与49个已知的铜中毒相关预后基因的共表达分析用于定义铜中毒相关的预后lncRNA。我们分别进行了LASSO算法和单变量和多变量Cox回归分析,以基于TCGA-LIHC数据集逐步确定铜中毒相关lncRNA的预后风险模型。随后,使用受试者操作特征(ROC)曲线、Kaplan-Meier生存曲线和预后列线图评估模型的预测性能。基因lncRNA与49个已知铜中毒相关基因的共表达分析在TCGA-LIHC数据集中鉴定了1359个铜中毒相关lncRNA。使用LASSO回归和Cox回归分析,用9种铜中毒相关的预后lncRNA(AC007998.3、AC003086.1、AC009974.2、IQCH-AS1、LINC0256 1、AC105345.1、ZFPM2-AS1、AL353708.1和WAC-AS1)构建预后模型。基于四个铜中毒相关lncRNA预后模型计算所有HCC患者样本的风险评分。所有HCC患者按照1:1的比例列分为高危和低危亚组。Kaplan-Meier生存曲线分析显示,高危组患者的总生存率(OS)显著低于低危组。主成分分析(PCA)证实,预后lncRNA模型准确区分了高危和低危HCC患者。此外,回归分析和ROC曲线证实了风险评分的预后价值。构建了一个包含风险评分和其他临床病理特征的列线图。列线图准确预测了HCC患者发生1年、3年和5年OS的概率。高风险患者的肿瘤突变负荷(TMB)评分高于低风险患者。与高危组相比,低风险组的HCC患者表现出更低的TIDE评分和更高的抗肿瘤药物敏感性。肝癌高危组和低危组的肿瘤免疫反应和肿瘤免疫细胞浸润有显著差异。我们的研究确定了一种9-铜相关lncRNA信号,该信号准确预测了HCC患者的预后、免疫治疗效果和肿瘤免疫微环境的状态。因此,这种铜中毒相关的lncRNA风险模型是HCC的一个潜在预后生物特征,在识别对免疫疗法有潜在反应的HCC患者方面显示出很高的临床价值。
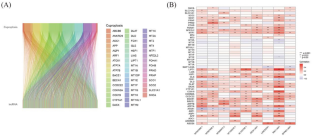
Cuproptosis is a novel programmed cell death pathway that is initiated by direct binding of copper to lipoylated tricarboxylic acid (TCA) cycle proteins. Recent studies have demonstrated that cuproptosis-related genes regulate tumorigenesis. However, the potential role and clinical significance of cuproptosis-related long noncoding RNAs (lncRNAs) in hepatocellular carcinoma (HCC) have not been established. We performed a bioinformatics analyses of RNA-sequencing data of HCC patients extracted from The Cancer Genome Atlas (TCGA) dataset to identify and validate a cuproptosis-related lncRNA prognostic signature. Furthermore, we analyzed the clinical significance of the prognostic signature of cuproptosis-related lncRNA in predicting the immunotherapeutic efficacy and the status of the tumor immune microenvironment. The RNA-sequencing data, genomic mutations, and clinical information were downloaded for 374 HCC samples and 50 normal liver samples from TCGA-Liver Hepatocellular Carcinoma (TCGA-LIHC) dataset. Co-expression analysis of Gene-lncRNA pairs with 49 known cuproptosis-related prognostic genes was used to define cuproptosis-related prognostic lncRNAs. We performed the LASSO algorithm and univariate and multivariate Cox regression analysis, respectively, to gradually identify the prognostic risk models of cuproptosis-related lncRNA based on the TCGA-LIHC dataset. Subsequently, the predictive performance of the model was evaluated using receiver operation characteristic (ROC) curves, Kaplan-Meier survival curves, and prognostic nomogram. The analysis of gene-lncRNA co-expression with 49 known cuproptosis-related genes identified 1359 cuproptosis-related lncRNAs in the TCGA-LIHC data set. A prognostic model was constructed with nine cuproptosis-related prognostic lncRNAs (AC007998.3, AC003086.1, AC009974.2, IQCH-AS1, LINC0256 1, AC105345.1, ZFPM2-AS1, AL353708.1 and WAC-AS1) using LASSO regression and Cox regression analyses. Risk scores were calculated for all HCC patient samples based on the four cuproptosis-related lncRNA prognostic models. All HCC patients were divided into high-risk and low-risk subgroups according to a 1:1 ratio column. The Kaplan-Meier survival curve analysis showed that the overall survival rate (OS) of the high-risk group patients was significantly lower than that of the low-risk group. The principal component analysis (PCA) confirmed that the prognostic lncRNA model accurately distinguished between high- and low-risk HCC patients. Furthermore, regression analysis as well as ROC curves confirmed the prognostic value of the risk score. A nomogram with risk scores and other clinicopathological characteristics was constructed. The nomogram accurately predicted the probability of 1-, 3-, and 5-year OS in HCC patients. Tumor mutation burden (TMB) scores were higher for high-risk patients than for low-risk patients. HCC patients in the low-risk group showed lower TIDE scores and greater sensitivity to antitumor drugs than those in the high-risk group. Tumor immune responses and tumor immune cell infiltration were significantly different between the high-risk and low-risk groups of patients with HCC. Our study identified a 9-cuproptosis-related lncRNA signature that accurately predicted prognosis, immunotherapeutic efficacy, and the status of the tumor immune microenvironment in HCC patients. Therefore, this cuproptosis-related lncRNA risk model is a potential prognostic biometric feature in HCC and shows high clinical value in identifying HCC patients who are potentially responsive to immunotherapy.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


