用于鉴定恶性疟原虫蛋白质组中药物-靶标相互作用的细胞热转移分析法
IF 16
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 57
摘要
尽管进行了数十年的研究,但人们对绝大多数抗疟药物的细胞靶点和作用模式知之甚少。我们最近证明,细胞热转移分析(CETSA)方案的两种变体:熔融曲线和等温剂量反应,代表了一种鉴定抗疟药物靶点的综合策略。CETSA 能够对作用机制未确定的未修饰抗疟药物进行全蛋白质组靶点筛选,提供药物与蛋白质直接相互作用的定量证据。实验工作流程包括用感兴趣的化合物处理恶性疟原虫感染的红细胞、加热使蛋白质变性、分离可溶性蛋白质、酶解、用串联质量标签标记肽、离线分馏和液相色谱-串联质谱分析。我们讨论了分析这种细胞内寄生虫所需的方法优化,包括富集寄生细胞和血红蛋白耗竭策略,以克服宿主红细胞中血红蛋白丰度高的问题。我们概述了使用 mineCETSA R 软件包进行有效数据处理的工作流程,该流程可为后续研究确定候选药物靶点的优先次序。整个方案可在两周内完成。作者展示了细胞热转移分析(CETSA)方案的两种变体--熔融曲线和等温剂量反应--用于完整细胞或裂解的恶性疟原虫样本,以鉴定抗疟药物靶点。本文章由计算机程序翻译,如有差异,请以英文原文为准。
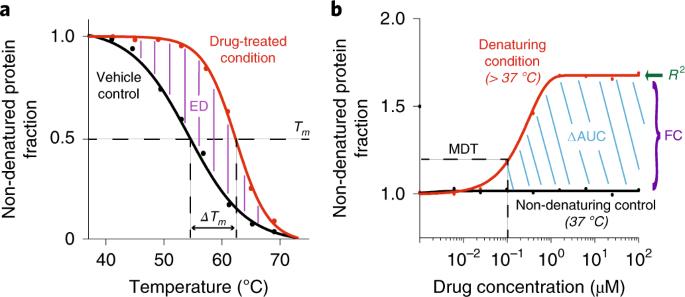
Cellular thermal shift assay for the identification of drug–target interactions in the Plasmodium falciparum proteome
Despite decades of research, little is known about the cellular targets and the mode of action of the vast majority of antimalarial drugs. We recently demonstrated that the cellular thermal shift assay (CETSA) protocol in its two variants: the melt curve and the isothermal dose-response, represents a comprehensive strategy for the identification of antimalarial drug targets. CETSA enables proteome-wide target screening for unmodified antimalarial compounds with undetermined mechanisms of action, providing quantitative evidence about direct drug–protein interactions. The experimental workflow involves treatment of P. falciparum–infected erythrocytes with a compound of interest, heat exposure to denature proteins, soluble protein isolation, enzymatic digestion, peptide labeling with tandem mass tags, offline fractionation, and liquid chromatography–tandem mass spectrometry analysis. Methodological optimizations necessary for the analysis of this intracellular parasite are discussed, including enrichment of parasitized cells and hemoglobin depletion strategies to overcome high hemoglobin abundance in the host red blood cells. We outline an effective data processing workflow using the mineCETSA R package, which enables prioritization of drug–target candidates for follow-up studies. The entire protocol can be completed within 2 weeks. The authors demonstrate two variants of the cellular thermal shift assay (CETSA) protocol—the melt curve and the isothermal dose–response—for use on intact cell or lysate P. falciparum samples for identification of antimalarial drug targets.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Nature Protocols
生物-生化研究方法
CiteScore
29.10
自引率
0.70%
发文量
128
审稿时长
4 months
期刊介绍:
Nature Protocols focuses on publishing protocols used to address significant biological and biomedical science research questions, including methods grounded in physics and chemistry with practical applications to biological problems. The journal caters to a primary audience of research scientists and, as such, exclusively publishes protocols with research applications. Protocols primarily aimed at influencing patient management and treatment decisions are not featured.
The specific techniques covered encompass a wide range, including but not limited to: Biochemistry, Cell biology, Cell culture, Chemical modification, Computational biology, Developmental biology, Epigenomics, Genetic analysis, Genetic modification, Genomics, Imaging, Immunology, Isolation, purification, and separation, Lipidomics, Metabolomics, Microbiology, Model organisms, Nanotechnology, Neuroscience, Nucleic-acid-based molecular biology, Pharmacology, Plant biology, Protein analysis, Proteomics, Spectroscopy, Structural biology, Synthetic chemistry, Tissue culture, Toxicology, and Virology.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


