景观规模的病毒分析揭示了乌兹别克斯坦丝绸之路枢纽的地方性和新兴蜜蜂病毒
IF 2.4
3区 生物学
Q1 ZOOLOGY
引用次数: 0
摘要
蜜蜂的健康正日益受到世界范围内复杂和不断演变的病毒景观的威胁;然而,尽管中亚处于历史和现代贸易路线的战略位置,但这方面的研究仍然不足。2024年,我们对乌兹别克斯坦11个地区32个城市的蜜蜂进行了全国性的病毒组调查,将14个汇集的RNA-seq文库的元基因组数据与RT-PCR验证和系统发育分析相结合。高质量测序平均每个池产生大约6000万次读取。我们从162个基因组序列(131个完整序列)中恢复了30种病毒,包括11种蜜蜂相关病毒和19种植物侵染病毒。所有样本均携带A型变形翼病毒(DWV-A),以合并感染DWV-B为主。我们的发现提供了第一个来自中亚的DWV-B全长基因组,揭示了它与欧洲菌株有97%的同源性。还检测到Sacbrood病毒(部分序列,约4.7 kb)和Lake Sinai病毒UZB的新变体。在乌兹别克斯坦首次对慢性蜜蜂麻痹病毒进行了完整的测序,发现varroa正黏液病毒-1表现出片段特异性分化。此外,我们还鉴定了两种新的植物病毒:Gulistan nepovirus 1和Arpa carmo-like virus 1。已查明的病毒的系统发育模式表明,乌兹别克斯坦是连接欧洲和亚洲病毒种群的遗传走廊。这些发现填补了关键的地理空白,强调了跨界监测的必要性,并为旨在保护传粉媒介和生态系统健康的未来诊断、流行病学和控制战略提供了基因组基线。本文章由计算机程序翻译,如有差异,请以英文原文为准。
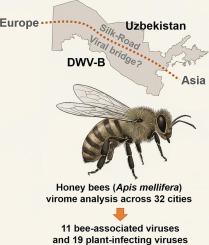
Landscape-scale virome analysis uncovers endemic and emerging honey bee viruses in the Silk-Road hub of Uzbekistan
Honey bee health is increasingly threatened worldwide by a complex and evolving viral landscape; however, this aspect in Central Asia remains understudied despite the region being strategically positioned along historic and modern trade routes. In 2024, we conducted a nationwide virome survey of Apis mellifera across 32 cities in 11 regions of Uzbekistan, combining the metagenomic data of 14 pooled RNA-seq libraries with RT-PCR validation and phylogenetic analyses. High-quality sequencing yielded an average of approximately 60 million reads per pool. We recovered 30 viral species from 162 genomic sequences (131 complete sequences), including 11 honey bee-associated and 19 plant-infecting viruses. All samples harbored deformed wing virus type A (DWV-A), and co-infection with DWV-B predominated. Our findings provided the first full-length DWV-B genomes from Central Asia, revealing that it had > 97 % identity to European strains. New variants of the Sacbrood virus (partial sequence, approximately 4.7 kb) and Lake Sinai virus UZB were also detected. The chronic bee paralysis virus was sequenced in full for the first time in Uzbekistan, and varroa orthomyxovirus-1 exhibited segment-specific divergence. Additionally, we identified two novel plant viruses: Gulistan nepovirus 1 and Arpa carmo-like virus 1. Phylogenetic patterns of the identified viruses indicate that Uzbekistan serves as a genetic corridor connecting European and Asian virus populations. These findings fill critical geographical gaps, underscore the need for transboundary surveillance, and provide a genomic baseline for future diagnostics, epidemiology, and control strategies aimed at safeguarding pollinator and ecosystem health.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
CiteScore
6.10
自引率
5.90%
发文量
94
审稿时长
1 months
期刊介绍:
The Journal of Invertebrate Pathology presents original research articles and notes on the induction and pathogenesis of diseases of invertebrates, including the suppression of diseases in beneficial species, and the use of diseases in controlling undesirable species. In addition, the journal publishes the results of physiological, morphological, genetic, immunological and ecological studies as related to the etiologic agents of diseases of invertebrates.
The Journal of Invertebrate Pathology is the adopted journal of the Society for Invertebrate Pathology, and is available to SIP members at a special reduced price.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


