阿尔茨海默病进展和认知恢复中的单细胞多区域表观基因组重接线
IF 42.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
阿尔茨海默病(AD)是一种以进行性认知能力下降为特征的神经退行性疾病,但其表观遗传基础仍然难以捉摸。在这里,我们生成并整合了来自111 AD 6个地区的384个死后大脑样本的350万个细胞的单细胞表观基因组学和转录组学图谱。我们确定了超过100万个候选顺式调控元件(cCREs),它们被组织成123个调控模块,跨越67个细胞亚型。我们定义了大规模的表观基因组区室和单细胞表观基因组信息,并描述了它们在阿尔茨海默病中的动态,揭示了阿尔茨海默病进展过程中广泛的表观基因组松弛和脑区域特异性和细胞类型特异性表观基因组侵蚀特征。这些表观基因组稳定性动态与细胞型比例变化、胶质细胞状态转变以及与阿尔茨海默病病理、认知障碍和认知恢复力相关的协调表观基因组和转录组失调密切相关。这项研究为阿尔茨海默病的进展和认知恢复提供了重要的见解,提出了一个全面的单细胞多组图谱,以促进对阿尔茨海默病的理解。本文章由计算机程序翻译,如有差异,请以英文原文为准。
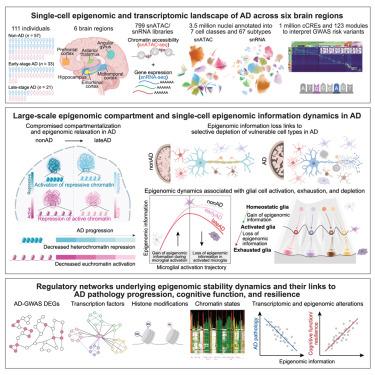
Single-cell multiregion epigenomic rewiring in Alzheimer’s disease progression and cognitive resilience
Alzheimer’s disease (AD) is a neurodegenerative disorder characterized by progressive cognitive decline, yet its epigenetic underpinnings remain elusive. Here, we generate and integrate single-cell epigenomic and transcriptomic profiles of 3.5 million cells from 384 postmortem brain samples across 6 regions in 111 AD and control individuals. We identify over 1 million candidate cis-regulatory elements (cCREs), organized into 123 regulatory modules across 67 cell subtypes. We define large-scale epigenomic compartments and single-cell epigenomic information and delineate their dynamics in AD, revealing widespread epigenome relaxation and brain-region-specific and cell-type-specific epigenomic erosion signatures during AD progression. These epigenomic stability dynamics are closely associated with cell-type proportion changes, glial cell-state transitions, and coordinated epigenomic and transcriptomic dysregulation linked to AD pathology, cognitive impairment, and cognitive resilience. This study provides critical insights into AD progression and cognitive resilience, presenting a comprehensive single-cell multiomic atlas to advance the understanding of AD.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Cell
生物-生化与分子生物学
CiteScore
110.00
自引率
0.80%
发文量
396
审稿时长
2 months
期刊介绍:
Cells is an international, peer-reviewed, open access journal that focuses on cell biology, molecular biology, and biophysics. It is affiliated with several societies, including the Spanish Society for Biochemistry and Molecular Biology (SEBBM), Nordic Autophagy Society (NAS), Spanish Society of Hematology and Hemotherapy (SEHH), and Society for Regenerative Medicine (Russian Federation) (RPO).
The journal publishes research findings of significant importance in various areas of experimental biology, such as cell biology, molecular biology, neuroscience, immunology, virology, microbiology, cancer, human genetics, systems biology, signaling, and disease mechanisms and therapeutics. The primary criterion for considering papers is whether the results contribute to significant conceptual advances or raise thought-provoking questions and hypotheses related to interesting and important biological inquiries.
In addition to primary research articles presented in four formats, Cells also features review and opinion articles in its "leading edge" section, discussing recent research advancements and topics of interest to its wide readership.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


