转移的进化途径
IF 66.8
1区 医学
Q1 ONCOLOGY
引用次数: 0
摘要
与早期癌变的生物学相比,人类肿瘤转移的进化尚不清楚。一个多世纪以来,临床医生和科学家们一直在争论,转移潜力是否是癌症的内在特性,由肿瘤创始细胞的分子特征预先决定,或者转移能力是否随着肿瘤的生长而逐步进化,类似于致癌改变的多阶段积累,导致第一个癌细胞的产生。在这个视角中,我研究了原发肿瘤和匹配转移的遗传分析如何区分这两种相互竞争的转移进化模型,特别强调了转移随机性的效用——一种定量测量,反映了转移是来自原发肿瘤亚克隆的随机选择,还是来自获得了促转移特征的特权谱系的后代。在具有高和低基线转移能力的肿瘤中可能的转移进化轨迹,以及在不同转移宿主部位的播种率和选择的作用进行了讨论。最后,我认为,如果我们共同努力,将现有的实验和理论工具应用于正确的患者群体,那么对人类转移生物学的开创性见解将立即成为可能。本文章由计算机程序翻译,如有差异,请以英文原文为准。
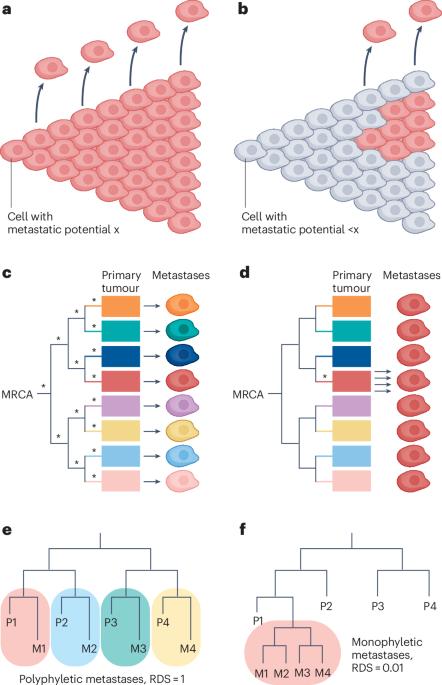

Evolutionary paths towards metastasis
The evolution of metastasis in humans is considerably less well understood than the biology of early carcinogenesis. For over a century, clinicians and scientists have been debating whether metastatic potential is the intrinsic property of a cancer, pre-determined by the molecular characteristics of the tumour founder cell, or whether metastatic capacity evolves in a stepwise fashion as the tumour grows, akin to the multistage accumulation of oncogenic alterations that give rise to the first cancer cell. In this Perspective, I examine how genetic analyses of primary tumours and matched metastases can distinguish between these two competing metastasis evolution models, with particular emphasis on the utility of metastatic randomness — a quantitative measure that reflects whether metastases arise from a random selection of primary tumour subclones or whether they are enriched for descendants of privileged lineages that have acquired pro-metastatic traits. Probable metastasis evolution trajectories in tumours with high and low baseline metastatic capacity are discussed, along with the role of seeding rates and selection at different metastatic host sites. Finally, I argue that trailblazing insights into human metastasis biology are immediately possible if we make a concerted effort to apply existing experimental and theoretical tools to the right patient cohorts. In this Perspective, Kamila Naxerova discusses how genetic analyses of primary tumours and matched metastases can distinguish between competing metastasis evolution models, arguing that further insights into human metastasis biology could be enabled by a framework that rigorously quantifies whether metastases descend from a nonrandom selection of primary tumour lineages.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Reviews Cancer
医学-肿瘤学
CiteScore
111.90
自引率
0.40%
发文量
97
审稿时长
6-12 weeks
期刊介绍:
Nature Reviews Cancer, a part of the Nature Reviews portfolio of journals, aims to be the premier source of reviews and commentaries for the scientific communities it serves. The correct abbreviation for abstracting and indexing purposes is Nat. Rev. Cancer. The international standard serial numbers (ISSN) for Nature Reviews Cancer are 1474-175X (print) and 1474-1768 (online). Unlike other journals, Nature Reviews Cancer does not have an external editorial board. Instead, all editorial decisions are made by a team of full-time professional editors who are PhD-level scientists. The journal publishes Research Highlights, Comments, Reviews, and Perspectives relevant to cancer researchers, ensuring that the articles reach the widest possible audience due to their broad scope.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


