增加结构稳定性和增强纳米体抗原结合的独特机制
IF 4.3
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
纳米体是单域抗体变体,证明自己是研究、疾病诊断和针对无数疾病因子的治疗方法的有力工具。然而,尽管有这种潜力,它们的顶楼呈现和结构稳定机制尚未得到充分探索。在这里,我们表明,与单克隆抗体不同,纳米体库通过在至少三个不同的方向上结合单个抗原,从而最大限度地采样抗原表面,这些方向与它们的伞形组成相关。一些纳米体的结构引导重组表明,在一个纳米体的框架3 (FR3)的高度保守区域的paratech或单点突变可以显著提高抗原亲和力,纳米体的稳定性,或两者兼有。相反,当“过度稳定”纳米体时,我们显示了对抗原亲和力的负面影响。总的来说,我们的研究结果提供了一种通用的策略,通过修改特定的残基来调节纳米体的亲和力,这些残基可以很容易地用于指导纳米体的优化和功能化。本文章由计算机程序翻译,如有差异,请以英文原文为准。
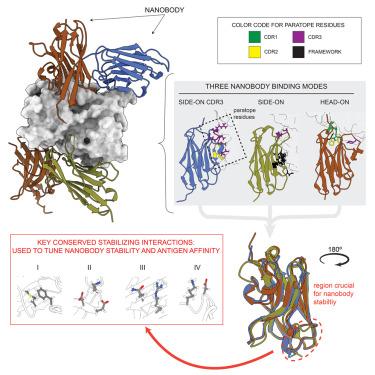
Unique mechanisms to increase structural stability and enhance antigen binding in nanobodies
Nanobodies are single domain antibody variants proving themselves to be compelling tools for research, disease diagnostics, and as therapeutics targeting a myriad of disease agents. However, despite this potential, their mechanisms of paratope presentation and structural stabilization have not been fully explored. Here, we show that unlike monoclonal antibodies, a nanobody repertoire maximizes sampling of an antigen surface by binding a single antigen in at least three different orientations, which are correlated with their paratope composition. Structure-guided reengineering of several nanobodies reveals that a single point mutation within the paratope or a highly conserved region of a nanobody’s framework 3 (FR3) can markedly improve antigen affinity, nanobody stability, or both. Conversely, we show the negative impact on antigen affinity when “over-stabilizing” nanobodies. Collectively our results provide a universal strategy to tune a nanobody’s affinity by modifying specific residues that can readily be applied to guide nanobody optimization and functionalization.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


