开发大型和复杂植物基因组的泛基因组及其表示格式
IF 13
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
摘要
泛基因组的发展通过捕获物种内完整的遗传多样性,彻底改变了基因组研究。泛基因组组装整合了来自多个个体的数据,构建了一个全面的基因组景观,揭示了核心和辅助基因组元素。这种方法能够识别新基因、结构变异和基因存在-缺失变异,为物种进化、适应和性状变异提供见解。表示泛基因组需要创新的可视化格式,有效地传达复杂的基因组结构和变异。目的综述了泛基因组的构建方法和最新进展,特别是植物基因组的构建。它研究了泛基因组表示的结构,包括格式比较、转换、可视化技术及其对加强作物改良策略的影响。先前的比较研究已经阐明了不同作物物种之间的新基因序列、拷贝数变异和存在-缺失变异。泛基因组的概念从广泛的基因型中捕获多种遗传变异,提供了一个物种基因组成的整体视角。然而,为具有更大基因组的植物构建泛基因组面临着挑战,包括管理大量基因组序列数据和理解种质内的遗传变异。为了应对这些挑战,研究人员已经探索了将物种多样性封装在一个称为泛基因组的单一组装中的成本效益替代方案。这包括减少基因组序列的数量,同时关注遗传变异。随着泛基因组概念在植物基因组学中的日益突出,一些软件工具已经出现,以促进泛基因组的构建。本文综述了植物泛基因组构建软件工具的开发与利用。它还讨论了适合下游分析的表示格式,为植物物种的遗传景观和进化动力学提供了有价值的见解。总之,这篇综述强调了泛基因组构建和表示格式在解决植物遗传结构方面的重要性,特别是那些具有复杂基因组的植物。它提供了最近进展的全面概述,有助于探索和理解植物遗传多样性。本文章由计算机程序翻译,如有差异,请以英文原文为准。
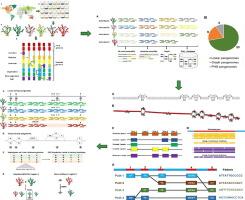
Developing pangenomes for large and complex plant genomes and their representation formats
Background
The development of pangenomes has revolutionized genomic studies by capturing the complete genetic diversity within a species. Pangenome assembly integrates data from multiple individuals to construct a comprehensive genomic landscape, revealing both core and accessory genomic elements. This approach enables the identification of novel genes, structural variations, and gene presence-absence variations, providing insights into species evolution, adaptation, and trait variation. Representing pangenomes requires innovative visualization formats that effectively convey the complex genomic structures and variations.Aim
This review delves into contemporary methodologies and recent advancements in constructing pangenomes, particularly in plant genomes. It examines the structure of pangenome representation, including format comparison, conversion, visualization techniques, and their implications for enhancing crop improvement strategies.Key scientific concepts of review
Earlier comparative studies have illuminated novel gene sequences, copy number variations, and presence-absence variations across diverse crop species. The concept of a pan-genome, which captures multiple genetic variations from a broad spectrum of genotypes, offers a holistic perspective of a species’ genetic makeup. However, constructing a pan-genome for plants with larger genomes poses challenges, including managing vast genome sequence data and comprehending the genetic variations within the germplasm. To address these challenges, researchers have explored cost-effective alternatives to encapsulate species diversity in a single assembly known as a pangenome. This involves reducing the volume of genome sequences while focusing on genetic variations. With the growing prominence of the pan-genome concept in plant genomics, several software tools have emerged to facilitate pangenome construction.This review sheds light on developing and utilizing software tools tailored for constructing pan-genomes in plants. It also discusses representation formats suitable for downstream analyses, offering valuable insights into the genetic landscape and evolutionary dynamics of plant species. In summary, this review underscores the significance of pan-genome construction and representation formats in resolving the genetic architecture of plants, particularly those with complex genomes. It provides a comprehensive overview of recent advancements, aiding in exploring and understanding plant genetic diversity.求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Journal of Advanced Research
Multidisciplinary-Multidisciplinary
CiteScore
21.60
自引率
0.90%
发文量
280
审稿时长
12 weeks
期刊介绍:
Journal of Advanced Research (J. Adv. Res.) is an applied/natural sciences, peer-reviewed journal that focuses on interdisciplinary research. The journal aims to contribute to applied research and knowledge worldwide through the publication of original and high-quality research articles in the fields of Medicine, Pharmaceutical Sciences, Dentistry, Physical Therapy, Veterinary Medicine, and Basic and Biological Sciences.
The following abstracting and indexing services cover the Journal of Advanced Research: PubMed/Medline, Essential Science Indicators, Web of Science, Scopus, PubMed Central, PubMed, Science Citation Index Expanded, Directory of Open Access Journals (DOAJ), and INSPEC.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


