真核细胞器动力学中涉及的Arf家族gtp酶的阿斯加德古菌起源
IF 20.5
1区 生物学
Q1 MICROBIOLOGY
引用次数: 0
摘要
真核生物的进化是生命史上的重大事件。与真核生物最接近的原核生物谱系是asgardarchaaeota,它们编码的蛋白质以前只在真核生物中发现,这为它们的古细菌祖先提供了深入了解。真核细胞以膜细胞器为特征,Arf家族gtpase通过在激活时将效应蛋白募集到膜上来调节细胞器动力学。Arf家族在真核生物中普遍存在,但其起源仍然难以捉摸。在这里,我们报道了一组广泛存在于asgardarchaaeota中的原核gtp酶,即arfr酶。系统发育分析显示真核Arf家族蛋白起源于ArfR组。代表性的asgardarchaaeta ArfR蛋白在酵母中的表达和x射线晶体学研究表明,ArfR GTPases具有膜结合机制和Arf家族蛋白特有的结构特征。我们的研究结果表明,Arf家族gtpase起源于真核生物的古细菌祖先,与真核生物早期进化的膜系统的各个方面一致。本文章由计算机程序翻译,如有差异,请以英文原文为准。

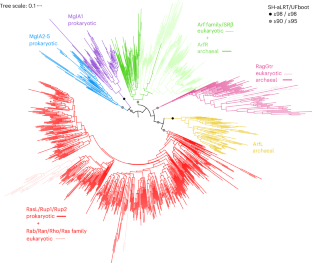
The Asgard archaeal origins of Arf family GTPases involved in eukaryotic organelle dynamics
The evolution of eukaryotes is a fundamental event in the history of life. The closest prokaryotic lineage to eukaryotes, the Asgardarchaeota, encode proteins previously found only in eukaryotes, providing insight into their archaeal ancestor. Eukaryotic cells are characterized by endomembrane organelles, and the Arf family GTPases regulate organelle dynamics by recruiting effector proteins to membranes upon activation. The Arf family is ubiquitous among eukaryotes, but its origins remain elusive. Here we report a group of prokaryotic GTPases, the ArfRs, which are widely present in Asgardarchaeota. Phylogenetic analyses reveal that eukaryotic Arf family proteins arose from the ArfR group. Expression of representative Asgardarchaeota ArfR proteins in yeast and X-ray crystallographic studies show that ArfR GTPases possess the mechanism of membrane binding and structural features unique to Arf family proteins. Our results indicate that Arf family GTPases originated in the archaeal ancestor of eukaryotes, consistent with aspects of the endomembrane system evolving early in eukaryogenesis. Eukaryotic Arf family proteins involved in endomembrane organelle dynamics arose from the Asgard archaeal ArfR GTPases, indicating this capability likely originated in the archaeal ancestor of eukaryotes.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Microbiology
Immunology and Microbiology-Microbiology
CiteScore
44.40
自引率
1.10%
发文量
226
期刊介绍:
Nature Microbiology aims to cover a comprehensive range of topics related to microorganisms. This includes:
Evolution: The journal is interested in exploring the evolutionary aspects of microorganisms. This may include research on their genetic diversity, adaptation, and speciation over time.
Physiology and cell biology: Nature Microbiology seeks to understand the functions and characteristics of microorganisms at the cellular and physiological levels. This may involve studying their metabolism, growth patterns, and cellular processes.
Interactions: The journal focuses on the interactions microorganisms have with each other, as well as their interactions with hosts or the environment. This encompasses investigations into microbial communities, symbiotic relationships, and microbial responses to different environments.
Societal significance: Nature Microbiology recognizes the societal impact of microorganisms and welcomes studies that explore their practical applications. This may include research on microbial diseases, biotechnology, or environmental remediation.
In summary, Nature Microbiology is interested in research related to the evolution, physiology and cell biology of microorganisms, their interactions, and their societal relevance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


