小鼠胚胎组织三维基因组和表观基因组的整合分析
IF 12.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
虽然最近在染色质可及性和组蛋白修饰模式的基础上,已经注释了一系列涉及小鼠胎儿发育的假定的顺式调控序列,但描述它们在发育调节基因表达中的作用仍然具有挑战性。为了填补这一空白,我们在7个小鼠胎儿组织和前脑的6个发育阶段绘制了基因启动子和远端序列之间的染色质接触图谱。我们确定了248,620个以14,138个蛋白质编码基因为中心的远程染色质相互作用,并表征了它们的组织间变异和发育动力学。将相互作用组与先前来自同一组织的表观基因组和转录组数据集进行综合分析,揭示了远端增强子的染色质接触和染色质状态之间的强相关性,以及预测靶基因的基因表达模式。我们预测了15098个候选增强子的靶基因,并用它们来注释人类基因组中含有人类疾病风险变体的同源候选增强子的靶基因。我们提供的证据表明,精神分裂症和其他成人疾病风险变异经常在胎儿增强子中发现,为成人疾病的胎儿起源假说提供了支持。本文章由计算机程序翻译,如有差异,请以英文原文为准。

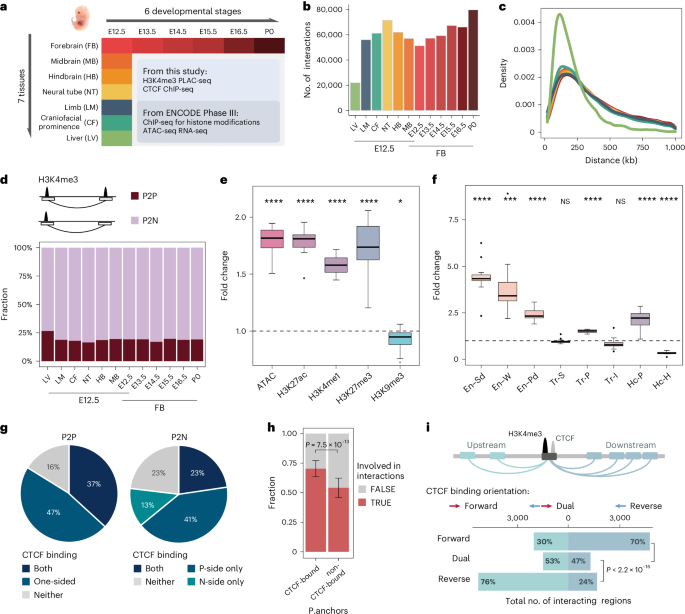
Integrative analysis of the 3D genome and epigenome in mouse embryonic tissues
While a rich set of putative cis-regulatory sequences involved in mouse fetal development have been annotated recently on the basis of chromatin accessibility and histone modification patterns, delineating their role in developmentally regulated gene expression continues to be challenging. To fill this gap, here we mapped chromatin contacts between gene promoters and distal sequences across the genome in seven mouse fetal tissues and across six developmental stages of the forebrain. We identified 248,620 long-range chromatin interactions centered at 14,138 protein-coding genes and characterized their tissue-to-tissue variations and developmental dynamics. Integrative analysis of the interactome with previous epigenome and transcriptome datasets from the same tissues revealed a strong correlation between the chromatin contacts and chromatin state at distal enhancers, as well as gene expression patterns at predicted target genes. We predicted target genes of 15,098 candidate enhancers and used them to annotate target genes of homologous candidate enhancers in the human genome that harbor risk variants of human diseases. We present evidence that schizophrenia and other adult disease risk variants are frequently found in fetal enhancers, providing support for the hypothesis of fetal origins of adult diseases. The authors here show that chromatin interactions during mouse fetal development are spatiotemporally dynamic. Integrating interactomes with other datasets predicts target genes for candidate enhancers and helps interpret noncoding risk variants in the human genome.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Structural & Molecular Biology
BIOCHEMISTRY & MOLECULAR BIOLOGY-BIOPHYSICS
CiteScore
22.00
自引率
1.80%
发文量
160
审稿时长
3-8 weeks
期刊介绍:
Nature Structural & Molecular Biology is a comprehensive platform that combines structural and molecular research. Our journal focuses on exploring the functional and mechanistic aspects of biological processes, emphasizing how molecular components collaborate to achieve a particular function. While structural data can shed light on these insights, our publication does not require them as a prerequisite.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


