用于功能性精准肿瘤学和发现科学的 PDX 模型
IF 66.8
1区 医学
Q1 ONCOLOGY
引用次数: 0
摘要
精准肿瘤学依赖于对不同肿瘤对各种疗法的反应进行详细的分子分析,目的是优化个体患者的治疗效果。患者衍生异种移植(PDX)模型一直是精准肿瘤学方法临床前验证的关键,它能够分析每种肿瘤的独特基因组情况,并测试根据特定突变、基因表达模式或信号异常预测有效的疗法。为了扩展这些标准的精准肿瘤学方法,该领域一直在努力用功能测定法(即功能精准肿瘤学 (FPO))来补充静态的、通常是描述性的测量方法。通过利用不同的 PDX 和 PDX 衍生模型,FPO 已成为一种有效的临床前和临床工具,利用体内和体外功能检测更精确地再现患者的生物学特性。在此,我们将探讨 PDX 和 PDX 衍生模型在精准肿瘤学和 FPO 方面的进展和局限性。我们还探讨了人工智能时代精准肿瘤学 PDX 模型的未来。整合这两个学科可能是快速、准确和经济有效的治疗预测的关键,从而彻底改变肿瘤学并为癌症患者提供最有效的个性化治疗。本文章由计算机程序翻译,如有差异,请以英文原文为准。

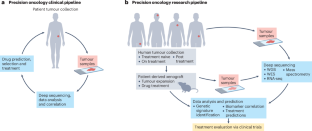
PDX models for functional precision oncology and discovery science
Precision oncology relies on detailed molecular analysis of how diverse tumours respond to various therapies, with the aim to optimize treatment outcomes for individual patients. Patient-derived xenograft (PDX) models have been key to preclinical validation of precision oncology approaches, enabling the analysis of each tumour’s unique genomic landscape and testing therapies that are predicted to be effective based on specific mutations, gene expression patterns or signalling abnormalities. To extend these standard precision oncology approaches, the field has strived to complement the otherwise static and often descriptive measurements with functional assays, termed functional precision oncology (FPO). By utilizing diverse PDX and PDX-derived models, FPO has gained traction as an effective preclinical and clinical tool to more precisely recapitulate patient biology using in vivo and ex vivo functional assays. Here, we explore advances and limitations of PDX and PDX-derived models for precision oncology and FPO. We also examine the future of PDX models for precision oncology in the age of artificial intelligence. Integrating these two disciplines could be the key to fast, accurate and cost-effective treatment prediction, revolutionizing oncology and providing patients with cancer with the most effective, personalized treatments. Patient-derived xenograft (PDX) models are valuable surrogates for drug testing in precision oncology. In this Review, Welm and colleagues outline the opportunities and challenges of PDX models, discuss their relevance in functional precision oncology for replicating patient biology and share their perspective on how integrating these models with artificial intelligence can enhance their utility for personalizing cancer treatment.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Reviews Cancer
医学-肿瘤学
CiteScore
111.90
自引率
0.40%
发文量
97
审稿时长
6-12 weeks
期刊介绍:
Nature Reviews Cancer, a part of the Nature Reviews portfolio of journals, aims to be the premier source of reviews and commentaries for the scientific communities it serves. The correct abbreviation for abstracting and indexing purposes is Nat. Rev. Cancer. The international standard serial numbers (ISSN) for Nature Reviews Cancer are 1474-175X (print) and 1474-1768 (online). Unlike other journals, Nature Reviews Cancer does not have an external editorial board. Instead, all editorial decisions are made by a team of full-time professional editors who are PhD-level scientists. The journal publishes Research Highlights, Comments, Reviews, and Perspectives relevant to cancer researchers, ensuring that the articles reach the widest possible audience due to their broad scope.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


