利用化学蛋白质组学分析完整蛋白质和原生蛋白质的新机遇
IF 5.7
2区 化学
Q1 CHEMISTRY, ANALYTICAL
引用次数: 0
摘要
本文章由计算机程序翻译,如有差异,请以英文原文为准。
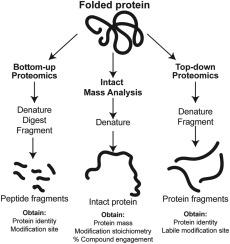
Emerging opportunities for intact and native protein analysis using chemical proteomics
Chemical proteomics has advanced small molecule ligand discovery by providing insights into protein-ligand binding mechanism and enabling medicinal chemistry optimization of protein selectivity on a global scale. Mass spectrometry is the predominant analytical method for chemoproteomics, and various approaches have been deployed to investigate and target a rapidly growing number of protein classes and biological systems. Two methods, intact mass analysis (IMA) and top-down proteomics (TDMS), have gained interest in recent years due to advancements in high resolution mass spectrometry instrumentation. Both methods apply mass spectrometry analysis at the proteoform level, as opposed to the peptide level of bottom-up proteomics (BUMS), thus addressing some of the challenges of protein inference and incomplete information on modification stoichiometry. This Review covers recent research progress utilizing MS-based proteomics methods, discussing in detail the capabilities and opportunities for improvement of each method. Further, heightened attention is given to IMA and TDMS, highlighting these methods’ strengths and considerations when utilized in chemoproteomic studies. Finally, we discuss the capabilities of native mass spectrometry (nMS) and how it can be used in chemoproteomics research to complement existing approaches to further advance the field of functional proteomics.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Analytica Chimica Acta
化学-分析化学
CiteScore
10.40
自引率
6.50%
发文量
1081
审稿时长
38 days
期刊介绍:
Analytica Chimica Acta has an open access mirror journal Analytica Chimica Acta: X, sharing the same aims and scope, editorial team, submission system and rigorous peer review.
Analytica Chimica Acta provides a forum for the rapid publication of original research, and critical, comprehensive reviews dealing with all aspects of fundamental and applied modern analytical chemistry. The journal welcomes the submission of research papers which report studies concerning the development of new and significant analytical methodologies. In determining the suitability of submitted articles for publication, particular scrutiny will be placed on the degree of novelty and impact of the research and the extent to which it adds to the existing body of knowledge in analytical chemistry.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


