肺炎患儿和健康对照组非类型流感嗜血杆菌基因组比较研究
IF 4.1
2区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
摘要
不可分型流感嗜血杆菌(NTHi)是引起呼吸道感染(包括儿童肺炎)的常见病原体,也可在无症状者的上呼吸道中发现。本研究探讨了健康儿童的 NTHi 菌株与急性或慢性社区获得性肺炎(CAP)患儿的 NTHi 菌株之间的基因组变异。通过细菌全基因组关联研究(bGWAS),我们对这些菌株进行了比较,以确定关键差异。我们的分析表明,约有 32% 的基因在共生和致病状态之间存在差异。值得注意的是,我们发现了肽聚糖生物合成途径的变化以及与肺炎相关的重要毒力因子。此外,我们还观察到健康组和肺炎组之间因 PBP3 突变而导致的对β-内酰胺类药物耐药性的显著差异,这一点已通过氨苄西林药敏试验得到证实,并以 D350N、S357N、S385T、L389F 突变模式为特征。这些发现有助于深入了解 NTHi 致病性的基因组基础,并为更有针对性的临床诊断和治疗提供依据。本文章由计算机程序翻译,如有差异,请以英文原文为准。
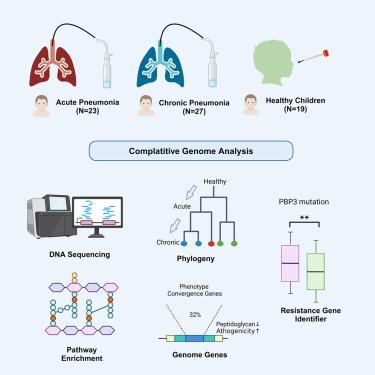
Comparative genomic study of non-typeable Haemophilus influenzae in children with pneumonia and healthy controls
Non-typeable Haemophilus influenzae (NTHi) is a common pathogen causing respiratory infections, including pneumonia in children, and can also be found in the upper respiratory tracts of asymptomatic individuals. This study examines genomic variations between NTHi strains from healthy children and those from children with acute or chronic community-acquired pneumonia (CAP). Using bacterial genome-wide association studies (bGWAS), we compared these strains to identify key differences. Our analysis revealed that approximately 32% of genes exhibit variations between commensal and pathogenic states. Notably, we identified changes in peptidoglycan biosynthesis pathways and significant virulence factors associated with pneumonia. Furthermore, we observed a significant difference in β-lactam resistance due to PBP3 mutations between the healthy and pneumonia groups, confirmed by the ampicillin susceptibility test and characterized by the mutation pattern D350N, S357N, S385T, L389F. These findings contribute valuable insights into the genomic basis of NTHi pathogenicity and may inform more targeted clinical diagnostics and treatments.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

iScience
Multidisciplinary-Multidisciplinary
CiteScore
7.20
自引率
1.70%
发文量
1972
审稿时长
6 weeks
期刊介绍:
Science has many big remaining questions. To address them, we will need to work collaboratively and across disciplines. The goal of iScience is to help fuel that type of interdisciplinary thinking. iScience is a new open-access journal from Cell Press that provides a platform for original research in the life, physical, and earth sciences. The primary criterion for publication in iScience is a significant contribution to a relevant field combined with robust results and underlying methodology. The advances appearing in iScience include both fundamental and applied investigations across this interdisciplinary range of topic areas. To support transparency in scientific investigation, we are happy to consider replication studies and papers that describe negative results.
We know you want your work to be published quickly and to be widely visible within your community and beyond. With the strong international reputation of Cell Press behind it, publication in iScience will help your work garner the attention and recognition it merits. Like all Cell Press journals, iScience prioritizes rapid publication. Our editorial team pays special attention to high-quality author service and to efficient, clear-cut decisions based on the information available within the manuscript. iScience taps into the expertise across Cell Press journals and selected partners to inform our editorial decisions and help publish your science in a timely and seamless way.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


