酵母基因组可在活细胞中全球访问
IF 12.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
真核生物的基因组被包装在染色质中,染色质由间隔规则的核小体凝聚成的丝状体组成,就像串在一起的珠子。核小体包含约 147 bp 的 DNA,几乎两次包裹在中央核心组蛋白八聚体周围。将 DNA 包入染色质对转录因子和其他需要访问其结合位点的蛋白质来说是一个挑战。因此,对 DNA 可及性的控制被认为在基因调控中起着关键作用。在这里,我们通过诱导 DNA 甲基转移酶的表达来测量活的芽殖酵母细胞基因组中 DNA 的可及性。我们发现,在活细胞中,基因组具有全局可及性,而在离体细胞核中则不同,在离体细胞核中,DNA 的可及性受到严重限制。基因体的甲基化速度仅略低于启动子,这表明酵母染色质在体内是高度动态的。相比之下,沉默基因座和中心粒受到强有力的保护。当细胞中依赖 ATP 的染色质重塑因子耗尽时,核小体位置会发生整体移动,这表明核小体的动态变化是这些酶之间竞争的结果。我们的结论是,染色质在活细胞中处于不断变化的状态,而在细胞核中则是静态的,这表明酵母中的DNA包装一般不是抑制性的。本文章由计算机程序翻译,如有差异,请以英文原文为准。

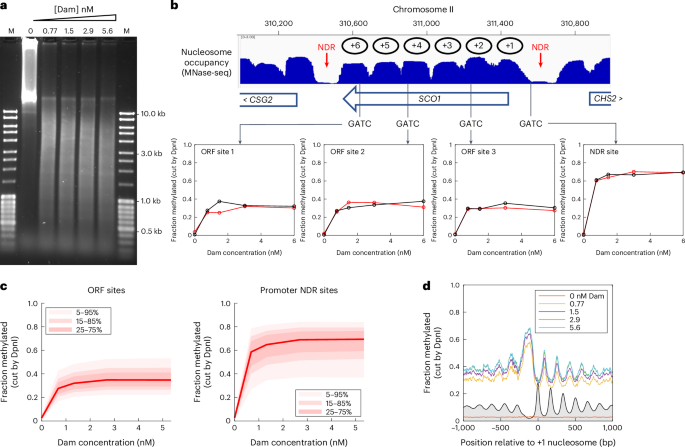
The yeast genome is globally accessible in living cells
Eukaryotic genomes are packaged into chromatin, which is composed of condensed filaments of regularly spaced nucleosomes, resembling beads on a string. The nucleosome contains ~147 bp of DNA wrapped almost twice around a central core histone octamer. The packaging of DNA into chromatin represents a challenge to transcription factors and other proteins requiring access to their binding sites. Consequently, control of DNA accessibility is thought to play a key role in gene regulation. Here we measure DNA accessibility genome wide in living budding yeast cells by inducible expression of DNA methyltransferases. We find that the genome is globally accessible in living cells, unlike in isolated nuclei, where DNA accessibility is severely restricted. Gene bodies are methylated at only slightly slower rates than promoters, indicating that yeast chromatin is highly dynamic in vivo. In contrast, silenced loci and centromeres are strongly protected. Global shifts in nucleosome positions occur in cells as they are depleted of ATP-dependent chromatin remodelers, suggesting that nucleosome dynamics result from competition among these enzymes. We conclude that chromatin is in a state of continuous flux in living cells, but static in nuclei, suggesting that DNA packaging in yeast is not generally repressive. The authors measure the accessibility of the yeast genome using DNA methylases. They show that the genome is globally accessible in living cells, except for centromeres and silenced loci, unlike in isolated nuclei, in which accessibility is strictly limited.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Structural & Molecular Biology
BIOCHEMISTRY & MOLECULAR BIOLOGY-BIOPHYSICS
CiteScore
22.00
自引率
1.80%
发文量
160
审稿时长
3-8 weeks
期刊介绍:
Nature Structural & Molecular Biology is a comprehensive platform that combines structural and molecular research. Our journal focuses on exploring the functional and mechanistic aspects of biological processes, emphasizing how molecular components collaborate to achieve a particular function. While structural data can shed light on these insights, our publication does not require them as a prerequisite.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


