转录组规模的 RNA 靶向 CRISPR 筛选揭示了人类细胞中的基本 lncRNA
IF 45.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
哺乳动物基因组承载着多种多样的 RNA,其中包括蛋白质编码和非编码转录本。然而,大多数长非编码 RNA(lncRNA)的功能作用仍然难以捉摸。通过使用 RNA 靶向 CRISPR-Cas13 筛选,我们在 5 个人类细胞系中探究了 6,200 个 lncRNA 的缺失如何影响细胞的适应性,并鉴定出 778 个具有特异性或广泛本质的 lncRNA。我们通过单个扰动确认了它们的基本性,并发现大多数基本的 lncRNA 是独立于其最近的蛋白编码基因运行的。通过对单细胞进行转录组分析,我们发现基本 lncRNAs 的缺失会影响细胞周期的进展并导致细胞凋亡。许多重要的lncRNA在发育过程中表现出跨组织的动态表达。利用 9,000 例原发性肿瘤,我们确定了那些在肿瘤中的表达与生存相关的 lncRNA,从而发现了新的生物标记物和潜在的治疗靶点。这项对功能性 lncRNA 的全转录组调查增进了我们对非编码转录本的了解,并展示了利用 Cas13 进行转录组规模非编码筛选的潜力。本文章由计算机程序翻译,如有差异,请以英文原文为准。
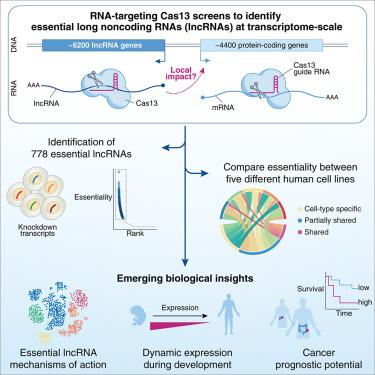
Transcriptome-scale RNA-targeting CRISPR screens reveal essential lncRNAs in human cells
Mammalian genomes host a diverse array of RNA that includes protein-coding and noncoding transcripts. However, the functional roles of most long noncoding RNAs (lncRNAs) remain elusive. Using RNA-targeting CRISPR-Cas13 screens, we probed how the loss of ∼6,200 lncRNAs impacts cell fitness across five human cell lines and identified 778 lncRNAs with context-specific or broad essentiality. We confirm their essentiality with individual perturbations and find that the majority of essential lncRNAs operate independently of their nearest protein-coding genes. Using transcriptome profiling in single cells, we discover that the loss of essential lncRNAs impairs cell-cycle progression and drives apoptosis. Many essential lncRNAs demonstrate dynamic expression across tissues during development. Using ∼9,000 primary tumors, we pinpoint those lncRNAs whose expression in tumors correlates with survival, yielding new biomarkers and potential therapeutic targets. This transcriptome-wide survey of functional lncRNAs advances our understanding of noncoding transcripts and demonstrates the potential of transcriptome-scale noncoding screens with Cas13.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Cell
生物-生化与分子生物学
CiteScore
110.00
自引率
0.80%
发文量
396
审稿时长
2 months
期刊介绍:
Cells is an international, peer-reviewed, open access journal that focuses on cell biology, molecular biology, and biophysics. It is affiliated with several societies, including the Spanish Society for Biochemistry and Molecular Biology (SEBBM), Nordic Autophagy Society (NAS), Spanish Society of Hematology and Hemotherapy (SEHH), and Society for Regenerative Medicine (Russian Federation) (RPO).
The journal publishes research findings of significant importance in various areas of experimental biology, such as cell biology, molecular biology, neuroscience, immunology, virology, microbiology, cancer, human genetics, systems biology, signaling, and disease mechanisms and therapeutics. The primary criterion for considering papers is whether the results contribute to significant conceptual advances or raise thought-provoking questions and hypotheses related to interesting and important biological inquiries.
In addition to primary research articles presented in four formats, Cells also features review and opinion articles in its "leading edge" section, discussing recent research advancements and topics of interest to its wide readership.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


