大肠杆菌细胞周期中染色体位点的三维定位和追踪。
IF 5.2
1区 生物学
Q1 BIOLOGY
引用次数: 0
摘要
基因在细胞内的位置可能会影响其表达,但目前还无法精确测量染色体基因座的三维位置。在二维空间中,可以使用与荧光蛋白融合的转录因子(TF)DNA 结合位点阵列来追踪基因座。然而,相同的二维数据可能来自不同的三维轨迹。在此,我们开发了一种深度学习方法,用于基于超分辨散光的活大肠杆菌细胞染色体基因座三维定位,在异质细胞背景下,信噪比约为 4,精度优于 61 nm。在确定染色体位点的空间定位时,我们发现有些位点在细胞周期的大部分时间里都位于核团的外围。对单个轨迹的分析表明,这些基因座在纵向(x)和径向(r)上都是亚扩散的,但单个基因座在微小时间尺度上探索了整个径向宽度。本文章由计算机程序翻译,如有差异,请以英文原文为准。
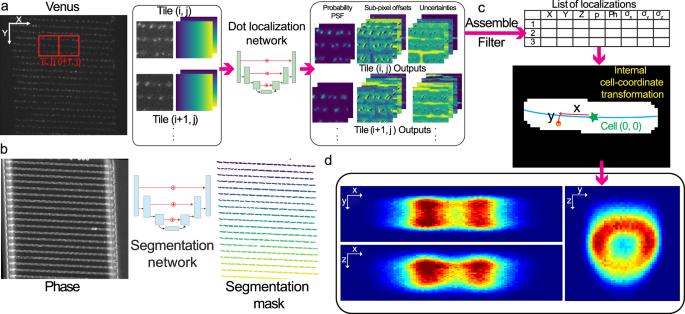
Three-dimensional localization and tracking of chromosomal loci throughout the Escherichia coli cell cycle
The intracellular position of genes may impact their expression, but it has not been possible to accurately measure the 3D position of chromosomal loci. In 2D, loci can be tracked using arrays of DNA-binding sites for transcription factors (TFs) fused with fluorescent proteins. However, the same 2D data can result from different 3D trajectories. Here, we have developed a deep learning method for super-resolved astigmatism-based 3D localization of chromosomal loci in live E. coli cells which enables a precision better than 61 nm at a signal-to-background ratio of ~4 on a heterogeneous cell background. Determining the spatial localization of chromosomal loci, we find that some loci are at the periphery of the nucleoid for large parts of the cell cycle. Analyses of individual trajectories reveal that these loci are subdiffusive both longitudinally (x) and radially (r), but that individual loci explore the full radial width on a minute time scale. The intracellular position of genes in the chromosome may impact their expression. To measure the 3D positions of chromosomal loci over the bacterial cell cycle, we have developed a combination of microfluidics, optics, and deep learning.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Communications Biology
Medicine-Medicine (miscellaneous)
CiteScore
8.60
自引率
1.70%
发文量
1233
审稿时长
13 weeks
期刊介绍:
Communications Biology is an open access journal from Nature Research publishing high-quality research, reviews and commentary in all areas of the biological sciences. Research papers published by the journal represent significant advances bringing new biological insight to a specialized area of research.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


