利用深度单瘤 RNA 测序探索人类躯体感觉的神经基础
IF 21.2
1区 医学
Q1 NEUROSCIENCES
引用次数: 0
摘要
躯体感觉的多变性来自异质性背根神经节(DRG)神经元。然而,由于技术上的困难,人类(h)DRG 神经元个体的体节转录组--解读其功能的关键信息--一直缺乏。在这项研究中,我们分离了单个 hDRG 神经元的体节,并进行了深度 RNA 测序(RNA-seq),平均每个神经元检测到超过 9,000 个独特基因,并确定了 16 种神经元类型。空间转录组学和 RNAscope 原位杂交证实并验证了这些结果。跨物种分析揭示了潜在痛觉神经元之间的差异,以及人类特异性神经元类型的可能存在。以分子谱为基础的微神经电图记录显示了人类感觉传入类型的温度感应特性。总之,通过采用单瘤深度RNA-seq和空间转录组学,我们生成了一个hDRG神经元图谱,为人类躯体感觉生理学提供了见解,并为转化工作奠定了基础。本文章由计算机程序翻译,如有差异,请以英文原文为准。
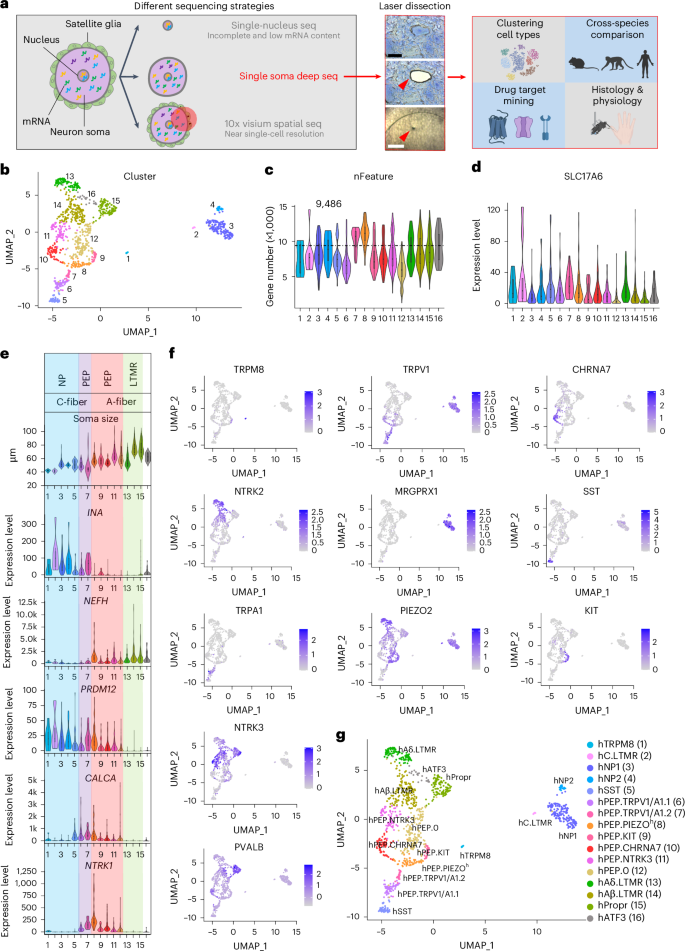
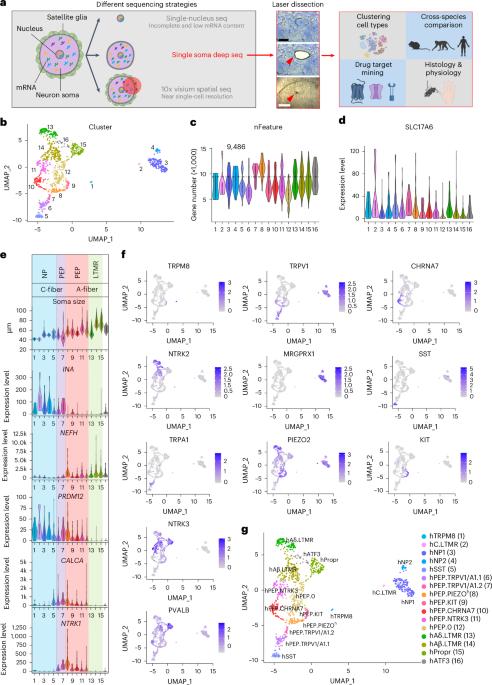
Leveraging deep single-soma RNA sequencing to explore the neural basis of human somatosensation
The versatility of somatosensation arises from heterogeneous dorsal root ganglion (DRG) neurons. However, soma transcriptomes of individual human (h)DRG neurons—critical information to decipher their functions—are lacking due to technical difficulties. In this study, we isolated somata from individual hDRG neurons and conducted deep RNA sequencing (RNA-seq) to detect, on average, over 9,000 unique genes per neuron, and we identified 16 neuronal types. These results were corroborated and validated by spatial transcriptomics and RNAscope in situ hybridization. Cross-species analyses revealed divergence among potential pain-sensing neurons and the likely existence of human-specific neuronal types. Molecular-profile-informed microneurography recordings revealed temperature-sensing properties across human sensory afferent types. In summary, by employing single-soma deep RNA-seq and spatial transcriptomics, we generated an hDRG neuron atlas, which provides insights into human somatosensory physiology and serves as a foundation for translational work. Dorsal root ganglia (DRGs) contain a plethora of neuron types. The authors show that the existence of human-specific DRG neuronal types and microneurography recordings reveal distinct temperature-sensing properties across human sensory afferent types.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature neuroscience
医学-神经科学
CiteScore
38.60
自引率
1.20%
发文量
212
审稿时长
1 months
期刊介绍:
Nature Neuroscience, a multidisciplinary journal, publishes papers of the utmost quality and significance across all realms of neuroscience. The editors welcome contributions spanning molecular, cellular, systems, and cognitive neuroscience, along with psychophysics, computational modeling, and nervous system disorders. While no area is off-limits, studies offering fundamental insights into nervous system function receive priority.
The journal offers high visibility to both readers and authors, fostering interdisciplinary communication and accessibility to a broad audience. It maintains high standards of copy editing and production, rigorous peer review, rapid publication, and operates independently from academic societies and other vested interests.
In addition to primary research, Nature Neuroscience features news and views, reviews, editorials, commentaries, perspectives, book reviews, and correspondence, aiming to serve as the voice of the global neuroscience community.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


