用于低温电子显微镜结构测定的 G 蛋白偶联受体复合物的原生质谱预筛选
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
G 蛋白偶联受体(GPCR)是一种重要的跨膜蛋白,在人类健康和疾病中发挥着关键作用。了解它们的原子级分子结构和构象状态对于推动药物开发至关重要。单颗粒低温电子显微镜(cryo-EM)的最新突破推动 GPCR 结构生物学进入了一个新时代。然而,由于 GPCR 的稳定性差且具有高动态特性,制备合适的 GPCR 样品及其复合物进行低温电子显微镜分析仍具有挑战性。在此,我们介绍了结合直接质谱技术(OBE-nMS+DMT)的在线缓冲液交换-原位质谱方法,该方法可促进高通量分析并指导样品制备。在冷冻电镜分析之前,我们采用这种方法对 GPR119-Gs 复合物样品进行了优化,结果在 200 kV Glacios 上仅采集了 396 个图像,就得到了 3.51 Å 分辨率的结构。这项研究表明,OBE-nMS+DMT 方法是预筛选 GPCR 和其他膜蛋白复合物冷冻电镜研究中样品条件的有力工具。本文章由计算机程序翻译,如有差异,请以英文原文为准。
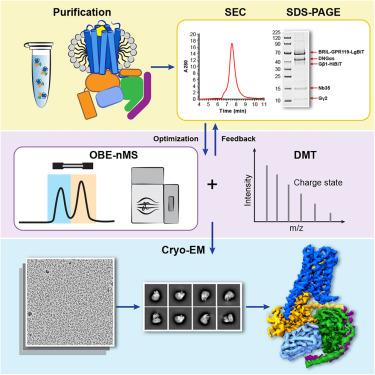
Native mass spectrometry prescreening of G protein-coupled receptor complexes for cryo-EM structure determination
G protein-coupled receptors (GPCRs) are essential transmembrane proteins playing key roles in human health and disease. Understanding their atomic-level molecular structure and conformational states is imperative for advancing drug development. Recent breakthroughs in single-particle cryogenic electron microscopy (cryo-EM) have propelled the structural biology of GPCRs into a new era. Nevertheless, the preparation of suitable GPCR samples and their complexes for cryo-EM analysis remains challenging due to their poor stability and highly dynamic nature. Here, we present our online buffer exchange-native MS method combined with Direct Mass Technology (OBE-nMS+DMT) which facilitates high-throughput analysis and guides sample preparation. We applied this method to optimize the GPR119-Gs complex sample prior to cryo-EM analysis, leading to a 3.51 Å resolution structure from only 396 movies collected on a 200 kV Glacios. This study suggests that the OBE-nMS+DMT method emerges as a powerful tool for prescreening sample conditions in cryo-EM studies of GPCRs and other membrane protein complexes.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


