过去五年药物与目标相互作用的研究进展。
IF 2.5
4区 生物学
Q2 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
摘要
药物与靶点相互作用(DTI)的鉴定是药物发现和药物再定位的重要步骤,在药物发现、药物再定位和再利用等多个领域具有很高的应用价值。然而,高昂的实验验证成本限制了其鉴定。相比之下,基于计算的方法既经济又高效。本综述首先综述了现有的化学基因组学方法,全面总结了用于预测 DTI 的流行数据库,并对近年来的特征编码进行了分类。随后概述并简要介绍了目前使用的 DTI 预测方法。然后详细讨论了最近五年(2020-2024 年)新提出的预测方法(包括基于网络表示学习和图神经网络的预测方法)的优缺点,评估了不同方法在各种数据集上的性能。最后,本综述探讨了未来 DTI 研究的潜在方向,强调如何通过结合大数据和新兴计算技术来提高预测的准确性和效率。本文章由计算机程序翻译,如有差异,请以英文原文为准。
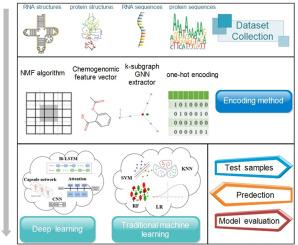
Research progress on Drug-Target Interactions in the last five years
The identification of Drug-Target Interaction (DTI) is an important step in drug discovery and drug repositioning, and has high application value in multiple fields such as drug discovery, drug repositioning, and repurposing. However, the high cost of experimental validation limits its identification. In contrast, computation-based approaches are both economical and efficient. This review first synthesizes existing chemical genomic approaches, provides a comprehensive summary of prevalent databases for predicting DTIs, and categorizes the feature encodings from recent years. This is followed by an overview and brief description of the methods currently in use for predicting DTIs. The strengths and weaknesses of newly proposed prediction methods in the last five years (2020–2024), including those based on network representation learning and graph neural networks, are then discussed in detail, evaluating the performance of the different methods on a wide range of datasets. Finally, this review explores potential directions for future DTI research, emphasizing how to improve prediction accuracy and efficiency by combining big data and emerging computing technologies.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Analytical biochemistry
生物-分析化学
CiteScore
5.70
自引率
0.00%
发文量
283
审稿时长
44 days
期刊介绍:
The journal''s title Analytical Biochemistry: Methods in the Biological Sciences declares its broad scope: methods for the basic biological sciences that include biochemistry, molecular genetics, cell biology, proteomics, immunology, bioinformatics and wherever the frontiers of research take the field.
The emphasis is on methods from the strictly analytical to the more preparative that would include novel approaches to protein purification as well as improvements in cell and organ culture. The actual techniques are equally inclusive ranging from aptamers to zymology.
The journal has been particularly active in:
-Analytical techniques for biological molecules-
Aptamer selection and utilization-
Biosensors-
Chromatography-
Cloning, sequencing and mutagenesis-
Electrochemical methods-
Electrophoresis-
Enzyme characterization methods-
Immunological approaches-
Mass spectrometry of proteins and nucleic acids-
Metabolomics-
Nano level techniques-
Optical spectroscopy in all its forms.
The journal is reluctant to include most drug and strictly clinical studies as there are more suitable publication platforms for these types of papers.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


