ANTIPASTI:利用正常模式和深度学习对抗体结合亲和力进行可解释的预测
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
抗体与其同源靶点的高结合亲和力是激发有效免疫反应以及将抗体用作研究和治疗工具的关键。在这里,我们提出了一种卷积神经网络模型 ANTIPASTI,该模型在预测抗体结合亲和力方面达到了最先进的水平,其输入数据是根据弹性网络模型得出的正态相关图表示的抗体-抗原结构。这种表征不仅能捕捉结构特征,还能捕捉局部和全局残基波动的能量模式。学习到的表征是可解释的:它们揭示了针对相同抗原类型的抗体之间结合模式的相似性,并可用于量化对结合亲和力有贡献的抗体区域的重要性。我们的研究结果表明了抗原印记在正常模式景观中的重要性,以及抗体区域之间的合作效应和长程相关性在决定结合亲和力方面的主导地位。本文章由计算机程序翻译,如有差异,请以英文原文为准。
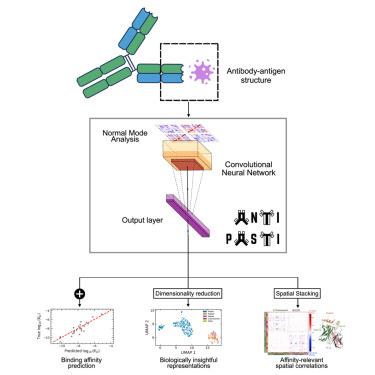
ANTIPASTI: Interpretable prediction of antibody binding affinity exploiting normal modes and deep learning
The high binding affinity of antibodies toward their cognate targets is key to eliciting effective immune responses, as well as to the use of antibodies as research and therapeutic tools. Here, we propose ANTIPASTI, a convolutional neural network model that achieves state-of-the-art performance in the prediction of antibody binding affinity using as input a representation of antibody-antigen structures in terms of normal mode correlation maps derived from elastic network models. This representation captures not only structural features but energetic patterns of local and global residue fluctuations. The learnt representations are interpretable: they reveal similarities of binding patterns among antibodies targeting the same antigen type, and can be used to quantify the importance of antibody regions contributing to binding affinity. Our results show the importance of the antigen imprint in the normal mode landscape, and the dominance of cooperative effects and long-range correlations between antibody regions to determine binding affinity.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


