喹啉类化合物可抑制作用于 DNA 的各种酶
IF 6.6
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
DNA 甲基化是一个关键的表观遗传过程,例如哺乳动物中的胞嘧啶-C5 甲基化和细菌中的腺嘌呤-N6 甲基化。开发非核苷类抑制剂来引起 DNA 低甲基化,对于治疗各种疾病而不产生现有的基于胞嘧啶的低甲基化药物的毒性至关重要。本研究鉴定了 15 种喹啉类类似物,特别是添加了甲胺(9)或甲基哌嗪(11)的化合物,它们对人类 DNMT1 和艰难梭菌 CamA 具有类似的低微摩尔抑制效力。这些化合物(9 和 11)通过小沟插层到与 CamA 结合的 DNA 中,引起构象转变,使催化结构域远离 DNA。这项研究增加了非核苷酸化合物通过 DNA 插层抑制 DNA 甲基转移酶的有限实例。此外,一些喹啉类似物还能抑制其他与 DNA 有相互作用的酶,如聚合酶和碱基切除修复糖基酶。最后,化合物 11 可通过激活癌细胞中的 p53 引起 DNA 损伤反应。本文章由计算机程序翻译,如有差异,请以英文原文为准。
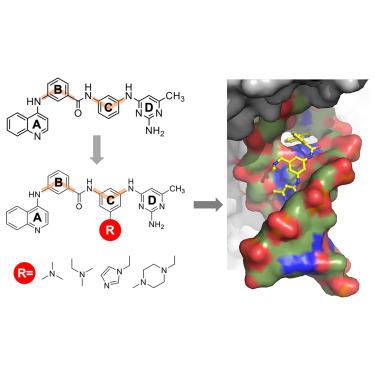

Quinoline-based compounds can inhibit diverse enzymes that act on DNA
DNA methylation, as exemplified by cytosine-C5 methylation in mammals and adenine-N6 methylation in bacteria, is a key epigenetic process. Developing non-nucleoside inhibitors to cause DNA hypomethylation is crucial for treating various conditions without the toxicities associated with existing cytidine-based hypomethylating agents. This study characterized fifteen quinoline-based analogs, particularly compounds with additions like a methylamine (9) or methylpiperazine (11), which demonstrate similar low micromolar inhibitory potency against human DNMT1 and Clostridioides difficile CamA. These compounds (9 and 11) intercalate into CamA-bound DNA via the minor groove, causing a conformational shift that moves the catalytic domain away from the DNA. This study adds to the limited examples of DNA methyltransferases being inhibited by non-nucleotide compounds through DNA intercalation. Additionally, some quinoline-based analogs inhibit other DNA-interacting enzymes, such as polymerases and base excision repair glycosylases. Finally, compound 11 elicits DNA damage response via p53 activation in cancer cells.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Cell Chemical Biology
Biochemistry, Genetics and Molecular Biology-Molecular Medicine
CiteScore
14.70
自引率
2.30%
发文量
143
期刊介绍:
Cell Chemical Biology, a Cell Press journal established in 1994 as Chemistry & Biology, focuses on publishing crucial advances in chemical biology research with broad appeal to our diverse community, spanning basic scientists to clinicians. Pioneering investigations at the chemistry-biology interface, the journal fosters collaboration between these disciplines. We encourage submissions providing significant conceptual advancements of broad interest across chemical, biological, clinical, and related fields. Particularly sought are articles utilizing chemical tools to perturb, visualize, and measure biological systems, offering unique insights into molecular mechanisms, disease biology, and therapeutics.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


