通过对短肽和 RNA 片段的原子模拟揭示核蛋白相分离亲和力
IF 4.8
2区 化学
Q2 CHEMISTRY, PHYSICAL
引用次数: 0
摘要
将蛋白质和核酸液-液相分离成凝聚相是确保细胞生物化学分区的一种多功能机制。RNA 分子在这些凝聚相中发挥着关键作用,特别是在转录调控和应激反应中,表现出多种热力学和动力学行为。然而,由于缩聚物的多组分和异质性质,破译支配蛋白质-RNA 缩聚物稳定性和动态的分子结构仍然具有挑战性。在这项研究中,我们对 20 种不同的混合物进行了原子模拟,这些混合物包含最小的 RNA 和肽片段,这使我们能够剖析所有 20 种氨基酸在 RNA 存在下的相分离亲和力。我们的研究结果阐明了协同促进或抑制蛋白质-RNA 相分离的化学特异性相互作用、水合曲线和离子效应。我们绘制了三元相互作用相图,确定了促进、维持、抑制和破坏蛋白质-RNA 簇的四组不同的残基。本文章由计算机程序翻译,如有差异,请以英文原文为准。
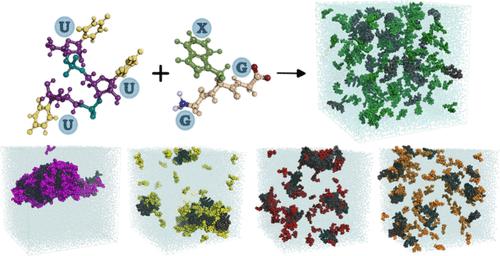
Nucleoprotein Phase-Separation Affinities Revealed via Atomistic Simulations of Short Peptide and RNA Fragments
Liquid–liquid phase separation of proteins and nucleic acids into condensate phases is a versatile mechanism for ensuring the compartmentalization of cellular biochemistry. RNA molecules play critical roles in these condensates, particularly in transcriptional regulation and stress responses, exhibiting a wide range of thermodynamic and dynamic behaviors. However, deciphering the molecular grammar that governs the stability and dynamics of protein–RNA condensates remains challenging due to the multicomponent and heterogeneous nature of condensates. In this study, we employ atomistic simulations of 20 distinct mixtures containing minimal RNA and peptide fragments which allows us to dissect the phase-separating affinities of all 20 amino acids in the presence of RNA. Our findings elucidate chemically specific interactions, hydration profiles, and ionic effects that synergistically promote or suppress protein–RNA phase separation. We map a ternary phase diagram of interactions, identifying four distinct groups of residues that promote, maintain, suppress, and disrupt protein–RNA clusters.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

The Journal of Physical Chemistry Letters
CHEMISTRY, PHYSICAL-NANOSCIENCE & NANOTECHNOLOGY
CiteScore
9.60
自引率
7.00%
发文量
1519
审稿时长
1.6 months
期刊介绍:
The Journal of Physical Chemistry (JPC) Letters is devoted to reporting new and original experimental and theoretical basic research of interest to physical chemists, biophysical chemists, chemical physicists, physicists, material scientists, and engineers. An important criterion for acceptance is that the paper reports a significant scientific advance and/or physical insight such that rapid publication is essential. Two issues of JPC Letters are published each month.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


