一类广泛分布的 C 尾锚定膜蛋白的核糖体失活作用
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
核糖体冬眠是一种常用策略,可在不利条件下保护核糖体并调节发育过程。在生命的所有领域中都发现了多种核糖体冬眠因子,但由于其结构的多样性和缺乏共同的失活机制,目前还不知道存在多少种不同的冬眠因子。在这里,我们发现 YqjD/ElaB/YgaM 旁系亲属最初是作为膜结合核糖体结合蛋白在大肠杆菌中被发现的,它们构成了一类丰富的核糖体冬眠蛋白,在所有蛋白细菌和其他一些细菌门中都是保守的。我们的数据表明,它们通过与 50S 核糖体亚基相互作用来抑制体外蛋白质合成。体内交联结合质谱分析表明,它们与核糖体隧道出口周围的蛋白质发生了特异性相互作用,甚至渗透到核糖体隧道中。因此,YqjD/ElaB/YgaM 通过阻断核糖体隧道抑制翻译,从而模拟抗菌肽和大环内酯类抗生素的活性。本文章由计算机程序翻译,如有差异,请以英文原文为准。
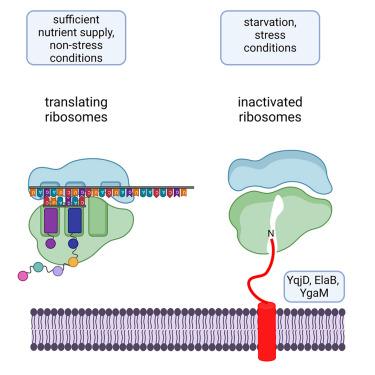
Ribosome-inactivation by a class of widely distributed C-tail anchored membrane proteins
Ribosome hibernation is a commonly used strategy that protects ribosomes under unfavorable conditions and regulates developmental processes. Multiple ribosome-hibernation factors have been identified in all domains of life, but due to their structural diversity and the lack of a common inactivation mechanism, it is currently unknown how many different hibernation factors exist. Here, we show that the YqjD/ElaB/YgaM paralogs, initially discovered as membrane-bound ribosome binding proteins in E. coli, constitute an abundant class of ribosome-hibernating proteins, which are conserved across all proteobacteria and some other bacterial phyla. Our data demonstrate that they inhibit in vitro protein synthesis by interacting with the 50S ribosomal subunit. In vivo cross-linking combined with mass spectrometry revealed their specific interactions with proteins surrounding the ribosomal tunnel exit and even their penetration into the ribosomal tunnel. Thus, YqjD/ElaB/YgaM inhibit translation by blocking the ribosomal tunnel and thus mimic the activity of antimicrobial peptides and macrolide antibiotics.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


