关于塑料化合物生物降解基因簇和途径的基因组学见解:揭示昆虫植物新陈代谢的多功能性 IITR165
IF 3.6
Q2 BIOTECHNOLOGY & APPLIED MICROBIOLOGY
引用次数: 0
摘要
研究人员对能够降解邻苯二甲酸二丁酯(DBP)、对苯二甲酸乙二醇酯(TPA)和聚对苯二甲酸乙二醇酯(PET)的 Dietzia kunjamensis IITR165 细菌进行了研究,以揭示其代谢途径。全基因组分析显示,该物种的环状染色体长达 3,477,711 bp,质粒长达 58,850 bp,GC 含量为 70.6%。在 3 311 个功能基因中,发现了邻苯二甲酸酯二氧合酶/脱羧酶(padAa1、padAb1、phtB、phtC)、烷烃单氧合酶(alkB)、邻苯二甲酸二烷基和单烷基水解酶以及二醇外二氧合酶。还发现了对苯二甲酸(tphA1A2A3 和 tphB)、苯甲酸(benABCD)和邻苯二酚(catABCD)的基因簇。根据 Q-TOF LC/MS-MS 分析,菌株 IITR165 在 96 小时内代谢了 1000 mg/L 的 TPA,半衰期为 15.36 h-1,代谢产物为邻苯二甲酸(PA)、苯甲酸(BA)和邻苯二酚。扫描电子显微镜照片显示了细菌处理后 PET 片材上广泛的生物膜和表面改性。研究发现了一种新型 PET 水解酶(PET165)蛋白,与已报道的 PET 酶有 45.70% 的氨基酸同源性,对接研究显示了与 PET 配体相互作用的保守催化三元组(丝氨酸-128、天冬氨酸-261 和组氨酸-287)。这种新型 PET 水解酶和 tpa 基因簇,以及参与尼龙和聚苯乙烯代谢的基因的存在,表明该细菌具有多功能性,可用于处理混合塑料污染的生态位。本文章由计算机程序翻译,如有差异,请以英文原文为准。
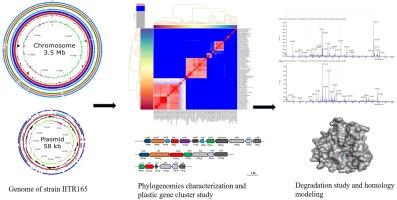
Genomic insights on gene clusters and pathways for the biodegradation of plastic compounds: Unravelling the metabolic versatility in a Dietzia kunjamensis IITR165
The Dietzia kunjamensis IITR165 bacterium, capable of degrading dibutyl phthalate (DBP), terephthalate (TPA), and polyethylene terephthalate (PET), was studied to uncover its metabolic pathways. Whole-genome analysis revealed a circular chromosome of 3,477,711 bp and a plasmid of 58,850 bp with 70.6 % GC content. Among 3,311 functional genes, phthalate dioxygenase/decarboxylase (padAa1, padAb1, phtB, phtC), alkane monooxygenase (alkB), di-and mono-alkyl phthalate hydrolase, and extra-diol dioxygenase were identified. Gene clusters for terephthalate (tphA1A2A3 and tphB), benzoic acid (benABCD), and catechol (catABCD) were also found. Strain IITR165 metabolized of 1000 mg/L of TPA in 96 h with a half-life of 15.36 h−1, producing phthalic acid (PA), benzoic acid (BA), and catechol as metabolites based on Q-TOF LC/MS-MS analysis. Scanning electron micrograph reveals the extensive biofilm and surface modification of PET sheet after bacterial treatment. A novel PET-hydrolase (PET165) protein, sharing 45.70 % amino acid homology with reported PETases, was discovered, with docking studies showing a conserved catalytic triad (Serine-128, Aspartate-261, and Histidine-287) interacting with the PET ligand. The presence of this novel PET hydrolase and the tpa gene cluster, along with genes involved in nylon, and polystyrene metabolism, indicates versatility of the bacterium useful in treatment of a mixed plastic contaminated ecological niches.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Current Research in Biotechnology
Biochemistry, Genetics and Molecular Biology-Biotechnology
CiteScore
6.70
自引率
3.60%
发文量
50
审稿时长
38 days
期刊介绍:
Current Research in Biotechnology (CRBIOT) is a new primary research, gold open access journal from Elsevier. CRBIOT publishes original papers, reviews, and short communications (including viewpoints and perspectives) resulting from research in biotechnology and biotech-associated disciplines.
Current Research in Biotechnology is a peer-reviewed gold open access (OA) journal and upon acceptance all articles are permanently and freely available. It is a companion to the highly regarded review journal Current Opinion in Biotechnology (2018 CiteScore 8.450) and is part of the Current Opinion and Research (CO+RE) suite of journals. All CO+RE journals leverage the Current Opinion legacy-of editorial excellence, high-impact, and global reach-to ensure they are a widely read resource that is integral to scientists' workflow.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


