IRF8 定义了出生后小胶质细胞的表观遗传景观,从而指导了它们的转录组程序
IF 27.7
1区 医学
Q1 IMMUNOLOGY
引用次数: 0
摘要
小胶质细胞是大脑中的先天性免疫细胞。转录因子 IRF8(干扰素调节因子 8)在小胶质细胞中高度表达。然而,它在出生后小胶质细胞发育过程中的作用尚不清楚。我们证明,IRF8与Sall1和PU.1一起逐步结合到出生后小胶质细胞的增强子区域,并在第14天后达到最大值。IRF8的结合与染色质可及性的逐步提高有关,而染色质可及性的提高先于小胶质细胞特异性转录组的启动。连续性和出生后Irf8缺失会导致小胶质细胞特性的丧失和疾病相关小胶质细胞(DAM)样基因的增加。单细胞(sc)RNA测序和单细胞转座酶可及染色质测序(scATAC-seq)的联合分析表明,染色质可及性和转录组在单细胞水平上存在相关性。小胶质细胞特异性DNA甲基化模式也需要IRF8。最后,在5xFAD模型中,组成型和出生后Irf8缺失会减少小胶质细胞与淀粉样β斑块的相互作用以及斑块的大小,从而减轻神经元的损失。总之,IRF8设定了表观遗传景观,这是出生后小胶质细胞基因表达所必需的。本文章由计算机程序翻译,如有差异,请以英文原文为准。
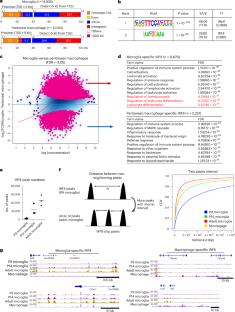
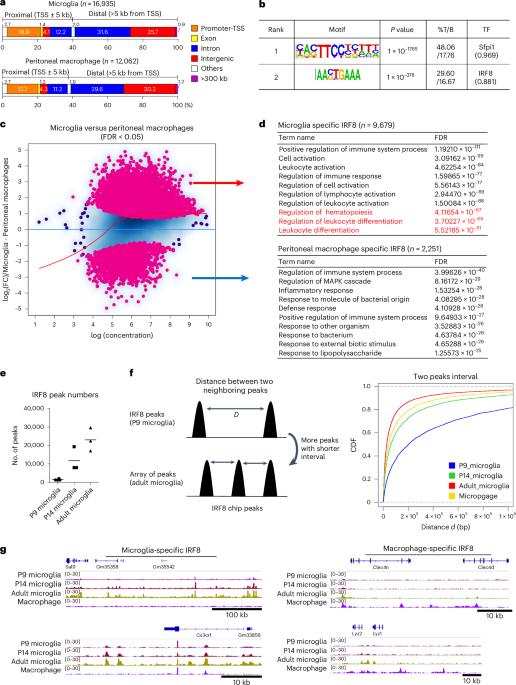
IRF8 defines the epigenetic landscape in postnatal microglia, thereby directing their transcriptome programs
Microglia are innate immune cells in the brain. Transcription factor IRF8 (interferon regulatory factor 8) is highly expressed in microglia. However, its role in postnatal microglia development is unknown. We demonstrate that IRF8 binds stepwise to enhancer regions of postnatal microglia along with Sall1 and PU.1, reaching a maximum after day 14. IRF8 binding correlated with a stepwise increase in chromatin accessibility, which preceded the initiation of microglia-specific transcriptome. Constitutive and postnatal Irf8 deletion led to a loss of microglia identity and gain of disease-associated microglia (DAM)-like genes. Combined analysis of single-cell (sc)RNA sequencing and single-cell transposase-accessible chromatin with sequencing (scATAC–seq) revealed a correlation between chromatin accessibility and transcriptome at a single-cell level. IRF8 was also required for microglia-specific DNA methylation patterns. Last, in the 5xFAD model, constitutive and postnatal Irf8 deletion reduced the interaction of microglia with amyloidβ plaques and the size of plaques, lessening neuronal loss. Together, IRF8 sets the epigenetic landscape, which is required for postnatal microglia gene expression. Saeki and colleagues show that IRF8 defines the epigenetic landscape in postnatal microglia, thereby directing their transcriptome programs.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Nature Immunology
医学-免疫学
CiteScore
40.00
自引率
2.30%
发文量
248
审稿时长
4-8 weeks
期刊介绍:
Nature Immunology is a monthly journal that publishes the highest quality research in all areas of immunology. The editorial decisions are made by a team of full-time professional editors. The journal prioritizes work that provides translational and/or fundamental insight into the workings of the immune system. It covers a wide range of topics including innate immunity and inflammation, development, immune receptors, signaling and apoptosis, antigen presentation, gene regulation and recombination, cellular and systemic immunity, vaccines, immune tolerance, autoimmunity, tumor immunology, and microbial immunopathology. In addition to publishing significant original research, Nature Immunology also includes comments, News and Views, research highlights, matters arising from readers, and reviews of the literature. The journal serves as a major conduit of top-quality information for the immunology community.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


