蛋白质组分析揭示早发型子痫前期患者胎盘组织中蛋白质表达的差异
IF 3.8
2区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
摘要
子痫前期是孕产妇和围产期发病率和死亡率的一个重要原因,但人们对其发病机制仍然知之甚少。本研究旨在通过对早发性子痫前期(EOPE)和正常妊娠胎盘中的差异蛋白进行比较分析,发现子痫前期的新型有效生物标记物。利用基于串联质量标签(TMT)的定量蛋白质组学,我们在15名EOPE患者和15名正常孕妇的胎盘组织中鉴定出了不同表达的蛋白质。这些蛋白质随后通过平行反应监测(PRM)进行了验证。我们的分析发现,与正常孕妇相比,EOPE 胎盘组织中共有 59 个差异表达蛋白,其中 25 个蛋白上调,34 个蛋白下调。通过PRM验证确认了6种蛋白的差异表达。我们的研究结果表明,这6种蛋白可能在EOPE的发病机制中起着关键作用,突出了雌激素信号通路和扩张型心肌病(DCM)通路在子痫前期发病中的潜在参与。这些数据已通过 iProX 合作伙伴存储库存入蛋白质组交换联盟(ProteomeXchange Consortium),标识符为 PXD055025。本文章由计算机程序翻译,如有差异,请以英文原文为准。
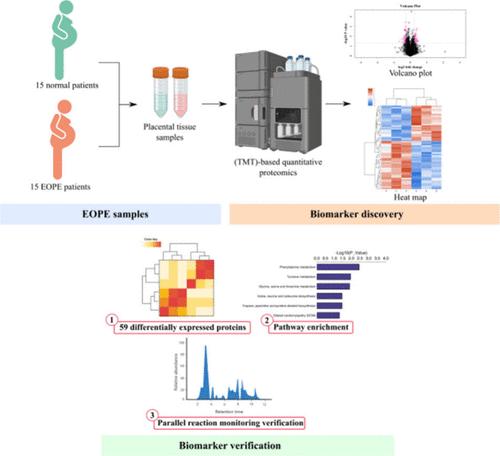
Proteomic Analysis Reveals Differential Protein Expression in Placental Tissues of Early-Onset Preeclampsia Patients
Preeclampsia, a significant cause of maternal and perinatal morbidity and mortality, remains poorly understood, in terms of its pathogenesis. This study aims to uncover novel and effective biomarkers for preeclampsia by conducting a comparative analysis of differential proteins in placentas from early onset preeclampsia (EOPE) and normal pregnancies. Utilizing tandem mass tag (TMT)-based quantitative proteomics, we identified differentially expressed proteins in placental tissues from 15 EOPE patients and 15 normal pregnant women. These proteins were subsequently validated by using parallel reaction monitoring (PRM). Our analysis revealed a total of 59 differentially expressed proteins, with 25 up-regulated and 34 down-regulated proteins in EOPE placental tissues compared to those from normal pregnancies. Validation through PRM confirmed the differential expression of 6 proteins. Our findings suggest these 6 proteins could play crucial roles in the pathogenesis of EOPE, highlighting the potential involvement of the estrogen signaling pathway and dilated cardiomyopathy (DCM) pathway in the development of preeclampsia. The data were deposited with the ProteomeXchange Consortium via the iProX partner repository with the identifier PXD055025.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Journal of Proteome Research
生物-生化研究方法
CiteScore
9.00
自引率
4.50%
发文量
251
审稿时长
3 months
期刊介绍:
Journal of Proteome Research publishes content encompassing all aspects of global protein analysis and function, including the dynamic aspects of genomics, spatio-temporal proteomics, metabonomics and metabolomics, clinical and agricultural proteomics, as well as advances in methodology including bioinformatics. The theme and emphasis is on a multidisciplinary approach to the life sciences through the synergy between the different types of "omics".
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


