肾小管上皮调节元件的变异介导了人类肾功能的大部分遗传差异
IF 29
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
摘要
肾功能衰竭是指肾功能下降到维持生命所需的临界值以下,是发病和死亡的主要原因。我们对英国生物库中的 406,504 人进行了全基因组关联研究(GWAS),确定了 430 个影响中年人肾功能的基因位点。为了研究受这些位点影响的细胞类型,我们将全基因组关联研究与利用单细胞转座酶可获取染色质测序(scATAC-seq)鉴定出的人类肾脏候选顺式调控元件(cCREs)进行了整合。总体而言,肾功能遗传性的 56% 定位于肾小管上皮细胞 cCREs,另有 7% 定位于肾荚膜细胞 cCREs。因此,成人肾功能的遗传差异大多是这两种细胞类型中基因表达改变的结果。通过增强子测定、等位基因特异性 scATAC-seq 和机器学习,我们发现许多肾功能变异会改变肾小管上皮细胞 cCRE 染色质的可及性和功能。利用 CRISPRi,我们确定了其中一些 cCRE 调控哪些基因,发现 NDRG1、CCNB1 和 STC1 与人类肾功能有关。本文章由计算机程序翻译,如有差异,请以英文原文为准。
Variants in tubule epithelial regulatory elements mediate most heritable differences in human kidney function
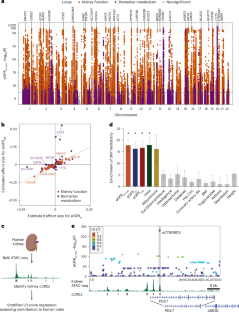
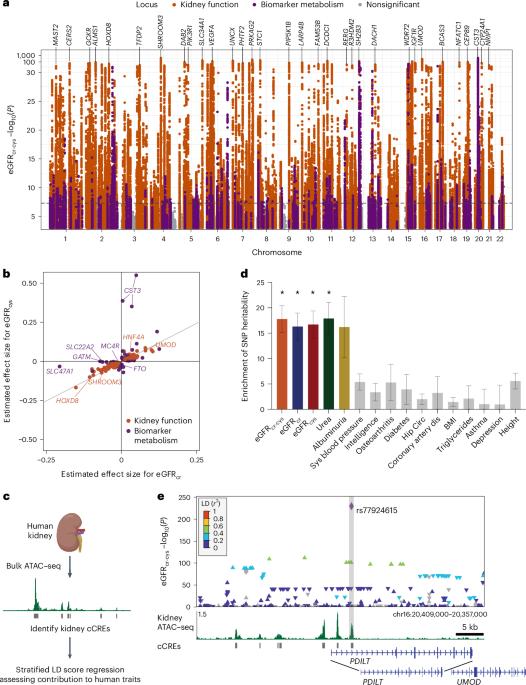
Kidney failure, the decrease of kidney function below a threshold necessary to support life, is a major cause of morbidity and mortality. We performed a genome-wide association study (GWAS) of 406,504 individuals in the UK Biobank, identifying 430 loci affecting kidney function in middle-aged adults. To investigate the cell types affected by these loci, we integrated the GWAS with human kidney candidate cis-regulatory elements (cCREs) identified using single-cell assay for transposase-accessible chromatin sequencing (scATAC–seq). Overall, 56% of kidney function heritability localized to kidney tubule epithelial cCREs and an additional 7% to kidney podocyte cCREs. Thus, most heritable differences in adult kidney function are a result of altered gene expression in these two cell types. Using enhancer assays, allele-specific scATAC–seq and machine learning, we found that many kidney function variants alter tubule epithelial cCRE chromatin accessibility and function. Using CRISPRi, we determined which genes some of these cCREs regulate, implicating NDRG1, CCNB1 and STC1 in human kidney function. Integration of genome-wide association analysis of 406,504 individuals in the UK Biobank and scATAC–seq data reveals that variants in tubule epithelial regulatory elements mediate most heritable differences in human kidney function.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:



