用于检测单细胞中 mRNA、细胞外蛋白质和细胞外蛋白质复合物的近距离测序。
IF 13.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
摘要
复杂的细胞功能是通过蛋白质复合物的协调形成和解离实现的。对信号配体的反应等功能可能包含数十种蛋白质和数百种复合物。直到最近,在单细胞水平测量多种蛋白质复合物还很困难。在这里,我们介绍一种逐步进行的近距离测序方法,它能同时测量蛋白质、mRNA 和位于细胞外膜上的数百种蛋白质复合物。我们指导用户完成探针创建、样品制备、染色、测序和蛋白质复合物的计算定量。例如,通过将转录测量与关联效应分子的测量结合起来,该方案使研究人员能够研究转录状态与细胞功能之间的相互作用,也可推广到其他配对事件。具有基础分子生物学和单细胞测序专业知识的用户只需花费大约 16 个小时,历时数天即可完成该方案。本文章由计算机程序翻译,如有差异,请以英文原文为准。

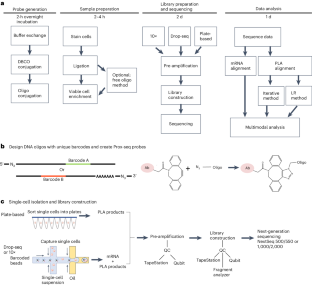
Proximity sequencing for the detection of mRNA, extracellular proteins and extracellular protein complexes in single cells
Complex cellular functions occur via the coordinated formation and dissociation of protein complexes. Functions such as the response to a signaling ligand can incorporate dozens of proteins and hundreds of complexes. Until recently, it has been difficult to measure multiple protein complexes at the single-cell level. Here, we present a step-by-step procedure for proximity sequencing, which enables the simultaneous measurement of proteins, mRNA and hundreds of protein complexes located on the outer membrane of cells. We guide the user through probe creation, sample preparation, staining, sequencing and computational quantification of protein complexes. This protocol empowers researchers to study, for example, the interplay between transcriptional states and cellular functions by coupling measurements of transcription to measurements of linked effector molecules, yet could be generalizable to other paired events. The protocol requires roughly 16 h spread over several days to complete by users with expertise in basic molecular biology and single-cell sequencing. Proximity sequencing uses a panel of antibodies to probe single cells for up to hundreds of targets simultaneously. The approach is capable of detecting targets that are located within 50–70 nm of each other and thus likely to form complexes.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Nature Protocols
生物-生化研究方法
CiteScore
29.10
自引率
0.70%
发文量
128
审稿时长
4 months
期刊介绍:
Nature Protocols focuses on publishing protocols used to address significant biological and biomedical science research questions, including methods grounded in physics and chemistry with practical applications to biological problems. The journal caters to a primary audience of research scientists and, as such, exclusively publishes protocols with research applications. Protocols primarily aimed at influencing patient management and treatment decisions are not featured.
The specific techniques covered encompass a wide range, including but not limited to: Biochemistry, Cell biology, Cell culture, Chemical modification, Computational biology, Developmental biology, Epigenomics, Genetic analysis, Genetic modification, Genomics, Imaging, Immunology, Isolation, purification, and separation, Lipidomics, Metabolomics, Microbiology, Model organisms, Nanotechnology, Neuroscience, Nucleic-acid-based molecular biology, Pharmacology, Plant biology, Protein analysis, Proteomics, Spectroscopy, Structural biology, Synthetic chemistry, Tissue culture, Toxicology, and Virology.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


