英格兰基因组学单倍型参考面板和英国生物数据库的估算
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
摘要
我们利用英格兰基因组学(Genomics England,GEL)数据集中的 78195 个个体建立了一个包含 3.42 亿个常染色体变异的参考面板,欧洲样本的相位切换错误率为 0.18%,英国白人样本中小等位基因频率低至 2 × 10-4 的变异的估算质量为 r2 = 0.75。GEL 估算的英国生物库全基因组关联分析确定了直接外显子组测序发现的关联的 70%(P < 2.18 × 10-11),同时将罕见变异的测试扩展到了整个基因组。编码变异在罕见变异全基因组关联结果中占主导地位,这意味着罕见非编码变异的破坏性影响较小。本文章由计算机程序翻译,如有差异,请以英文原文为准。
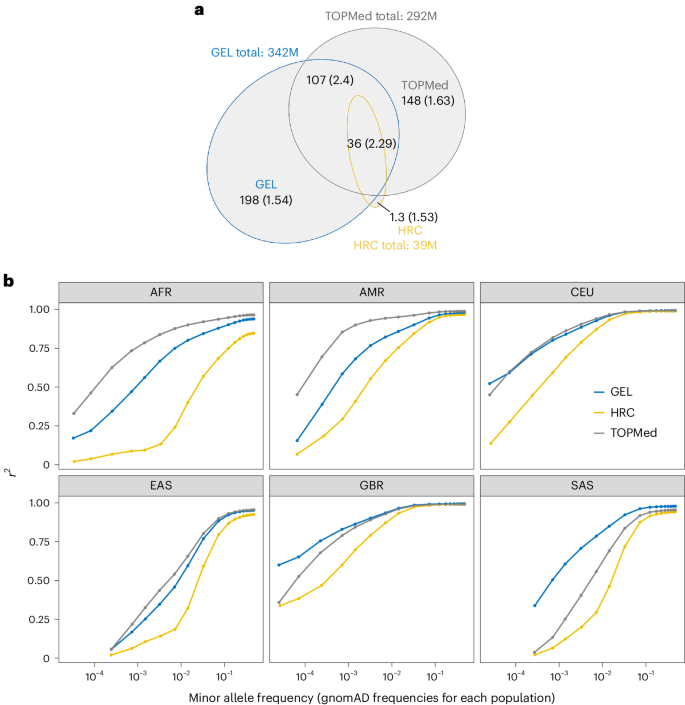
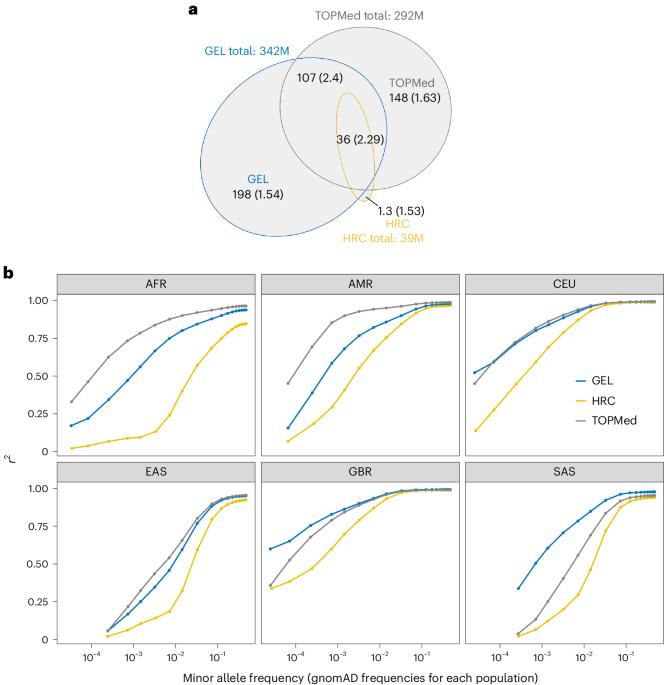
A Genomics England haplotype reference panel and imputation of UK Biobank
We built a reference panel with 342 million autosomal variants using 78,195 individuals from the Genomics England (GEL) dataset, achieving a phasing switch error rate of 0.18% for European samples and imputation quality of r2 = 0.75 for variants with minor allele frequencies as low as 2 × 10−4 in white British samples. The GEL-imputed UK Biobank genome-wide association analysis identified 70% of associations found by direct exome sequencing (P < 2.18 × 10−11), while extending testing of rare variants to the entire genome. Coding variants dominated the rare-variant genome-wide association results, implying less disruptive effects of rare non-coding variants. A Genomics England haplotype reference panel constructed using sequence data from 78,195 individuals improves phasing and imputation accuracy. Imputation of the UK Biobank using this panel enables genome-wide rare-variant association analyses.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


