用序列到功能模型揭示基因调控。
IF 36.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
摘要
序列到功能模型利用现代人工智能和大规模功能基因组数据集的最新进展,学习基因组 DNA 与其多层基因调控功能之间的关系。这些模型有望揭示跨细胞生物学各层次的机理关系,从而改变我们对顺式基因调控的理解,为发现疾病机理开辟新途径。本文章由计算机程序翻译,如有差异,请以英文原文为准。
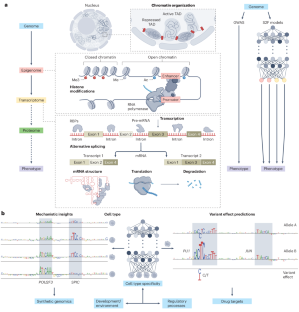
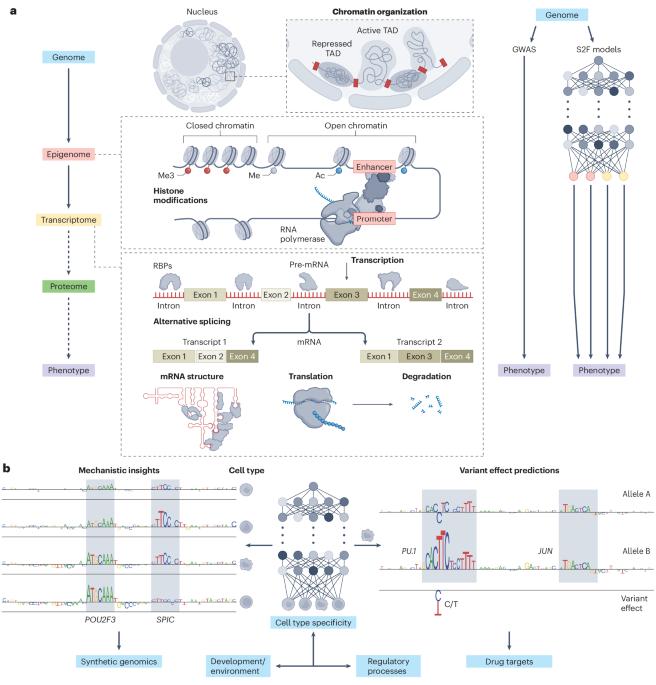
Unlocking gene regulation with sequence-to-function models
By exploiting recent advances in modern artificial intelligence and large-scale functional genomic datasets, sequence-to-function models learn the relationship between genomic DNA and its multilayer gene regulatory functions. These models are poised to uncover mechanistic relationships across layers of cellular biology, which will transform our understanding of cis gene regulation and open new avenues for discovering disease mechanisms.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Methods
生物-生化研究方法
CiteScore
58.70
自引率
1.70%
发文量
326
审稿时长
1 months
期刊介绍:
Nature Methods is a monthly journal that focuses on publishing innovative methods and substantial enhancements to fundamental life sciences research techniques. Geared towards a diverse, interdisciplinary readership of researchers in academia and industry engaged in laboratory work, the journal offers new tools for research and emphasizes the immediate practical significance of the featured work. It publishes primary research papers and reviews recent technical and methodological advancements, with a particular interest in primary methods papers relevant to the biological and biomedical sciences. This includes methods rooted in chemistry with practical applications for studying biological problems.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


