水稻胚乳中持久性和阶段特异性印迹基因的多层表观遗传控制
IF 15.8
1区 生物学
Q1 PLANT SCIENCES
引用次数: 0
摘要
在被子植物中,基因组印记的表观遗传图谱在受精前就已建立。然而,表观遗传修饰与印记表达之间的因果关系尚未完全明了。在这项研究中,我们根据对水稻(Oryza sativa)胚乳的时程转录组分析,对 "持久性 "和 "阶段特异性 "印记基因进行了分类,并将它们与单个时间点的表观遗传修饰进行了比较。阶段特异性印记基因的表观遗传修饰水平相对较低,而持久性印记基因的表观遗传修饰水平则要高得多。总体趋势显示,母系表达的印记基因的母系等位基因被DNA去甲基化激活,而具有基因体甲基化(gbM)的父系表达的印记基因的母系等位基因则被DNA去甲基化和H3K27me3沉积沉默,这些区域与Tc/Mar-Stowaway相关的富集基序有关。我们的发现有助于深入了解基因组印记的稳定性以及与胚乳发育、不同细胞类型和亲本基因型相关的潜在变化。本文章由计算机程序翻译,如有差异,请以英文原文为准。
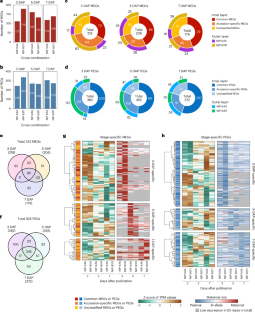
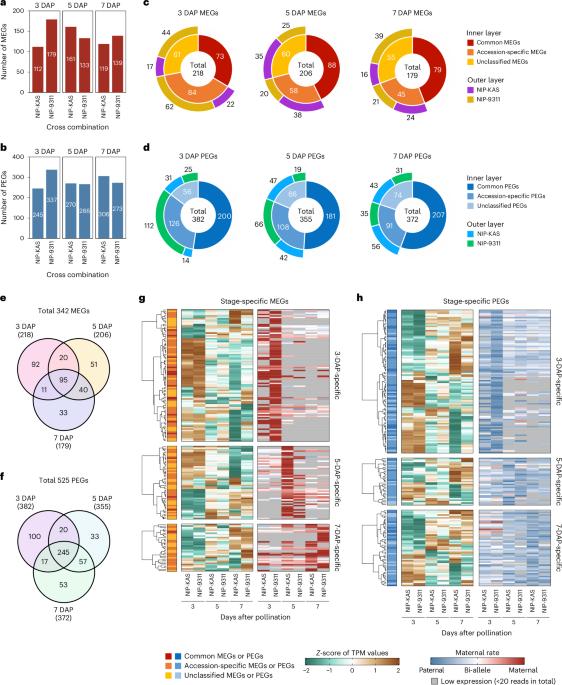
Multilayered epigenetic control of persistent and stage-specific imprinted genes in rice endosperm
In angiosperms, epigenetic profiles for genomic imprinting are established before fertilization. However, the causal relationships between epigenetic modifications and imprinted expression are not fully understood. In this study, we classified ‘persistent’ and ‘stage-specific’ imprinted genes on the basis of time-course transcriptome analysis in rice (Oryza sativa) endosperm and compared them to epigenetic modifications at a single time point. While the levels of epigenetic modifications are relatively low in stage-specific imprinted genes, they are considerably higher in persistent imprinted genes. Overall trends revealed that the maternal alleles of maternally expressed imprinted genes are activated by DNA demethylation, while the maternal alleles of paternally expressed imprinted genes with gene body methylation (gbM) are silenced by DNA demethylation and H3K27me3 deposition, and these regions are associated with an enriched motif related to Tc/Mar-Stowaway. Our findings provide insight into the stability of genomic imprinting and the potential variations associated with endosperm development, different cell types and parental genotypes. Time-course analysis of rice imprinted genes revealed persistent and stage-specific imprinted genes. These two classes showed differences in the degree of epigenetic status and biased expression in a temporal and spatial manner in the endosperm.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Nature Plants
PLANT SCIENCES-
CiteScore
25.30
自引率
2.20%
发文量
196
期刊介绍:
Nature Plants is an online-only, monthly journal publishing the best research on plants — from their evolution, development, metabolism and environmental interactions to their societal significance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


