模板辅助共价修饰是共价分子胶活性的基础
IF 12.9
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
分子粘合剂是一种诱导接近的小分子,已成为一种极具吸引力的治疗方法。然而,分子胶的开发仍然充满挑战,需要创新的机制策略来稳定新蛋白界面并加速发现。在这里,我们揭示了一种称为 "模板辅助共价修饰 "的反式标记共价分子胶合机制。我们发现了一系列新的 BRD4 分子胶降解剂,它们能将 CUL4DCAF16 连接酶招募到 BRD4 的第二个溴结构域(BRD4BD2)。通过全面的生化、结构和诱变分析,我们阐明了 DCAF16 和 BRD4BD2 之间预先存在的结构互补性如何作为模板,优化降解器的方向,对 DCAF16Cys58 进行共价修饰。这一过程稳定了BRD4-降解剂-DCAF16三元复合物的形成,促进了BRD4的降解。我们发现,一个降解子集也能通过反式标记 GAK 来诱导 GAK-BRD4BD2 相互作用,这证明了降解的普遍性。总之,我们的工作确立了 "模板辅助共价修饰 "作为共价分子胶合剂的一种机制,为近距离驱动药理学开辟了一条新路。本文章由计算机程序翻译,如有差异,请以英文原文为准。

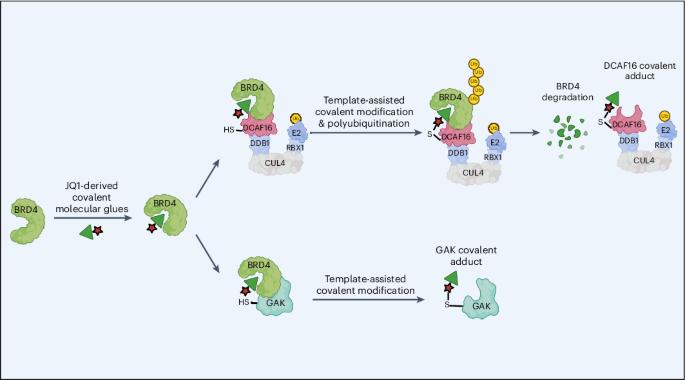
Template-assisted covalent modification underlies activity of covalent molecular glues
Molecular glues are proximity-inducing small molecules that have emerged as an attractive therapeutic approach. However, developing molecular glues remains challenging, requiring innovative mechanistic strategies to stabilize neoprotein interfaces and expedite discovery. Here we unveil a trans-labeling covalent molecular glue mechanism, termed ‘template-assisted covalent modification’. We identified a new series of BRD4 molecular glue degraders that recruit CUL4DCAF16 ligase to the second bromodomain of BRD4 (BRD4BD2). Through comprehensive biochemical, structural and mutagenesis analyses, we elucidated how pre-existing structural complementarity between DCAF16 and BRD4BD2 serves as a template to optimally orient the degrader for covalent modification of DCAF16Cys58. This process stabilizes the formation of BRD4–degrader–DCAF16 ternary complex and facilitates BRD4 degradation. Supporting generalizability, we found that a subset of degraders also induces GAK–BRD4BD2 interaction through trans-labeling of GAK. Together, our work establishes ‘template-assisted covalent modification’ as a mechanism for covalent molecular glues, which opens a new path to proximity-driven pharmacology. Characterization of DCAF16-based BRD4 molecular glue degraders revealed a trans-labeling mechanism termed ‘template-assisted covalent modification’, which opens a new path for proximity-driven pharmacology.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature chemical biology
生物-生化与分子生物学
CiteScore
23.90
自引率
1.40%
发文量
238
审稿时长
12 months
期刊介绍:
Nature Chemical Biology stands as an esteemed international monthly journal, offering a prominent platform for the chemical biology community to showcase top-tier original research and commentary. Operating at the crossroads of chemistry, biology, and related disciplines, chemical biology utilizes scientific ideas and approaches to comprehend and manipulate biological systems with molecular precision.
The journal embraces contributions from the growing community of chemical biologists, encompassing insights from chemists applying principles and tools to biological inquiries and biologists striving to comprehend and control molecular-level biological processes. We prioritize studies unveiling significant conceptual or practical advancements in areas where chemistry and biology intersect, emphasizing basic research, especially those reporting novel chemical or biological tools and offering profound molecular-level insights into underlying biological mechanisms.
Nature Chemical Biology also welcomes manuscripts describing applied molecular studies at the chemistry-biology interface due to the broad utility of chemical biology approaches in manipulating or engineering biological systems. Irrespective of scientific focus, we actively seek submissions that creatively blend chemistry and biology, particularly those providing substantial conceptual or methodological breakthroughs with the potential to open innovative research avenues. The journal maintains a robust and impartial review process, emphasizing thorough chemical and biological characterization.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


