定位的 PABP-RNA 中枢在发育过程中实现了信号依赖性 mRNA 衰减。
IF 12.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
信号通路主要通过改变基因表达来驱动细胞命运的转变。然而,对信号线索做出快速和选择性转录组重配的机制仍然难以捉摸。在这里,我们利用深度学习来解构引发细胞外信号调节激酶(ERK)-介导原活化蛋白激酶激酶(MEK)诱导的幼稚多能性mRNA衰变的序列决定因素和反式作用调节因子。通过ERK-MEK磷酸化LIN28A,使pLIN28A重新定位到幼稚多能性mRNA高度富含A+U的3'非翻译区(3'UTR)末端,衰变的时间与胚胎植入相关联。有趣的是,这些富含 A+U 的 3'UTR 末端是多聚(A)结合蛋白(PABP)的结合枢纽,可在信号诱导下与 LIN28A 聚合。AUU基序的多价性决定了pLIN28A-PABP汇聚的有效性,它能增强PABP 3'UTR的结合,减少对poly(A)尾的保护,激活mRNA的衰变,使其向原始多能性发展。因此,信号诱导的 LIN28A 与 PABP-RNA 中枢的聚合推动了对幼稚 mRNA 的快速衰变选择,实现了转录组的重塑,从而确保了快速的发育进程。本文章由计算机程序翻译,如有差异,请以英文原文为准。
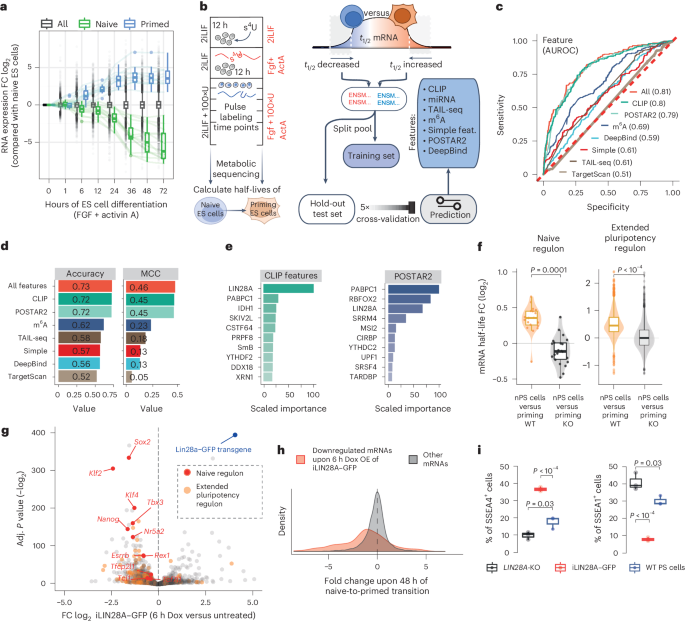
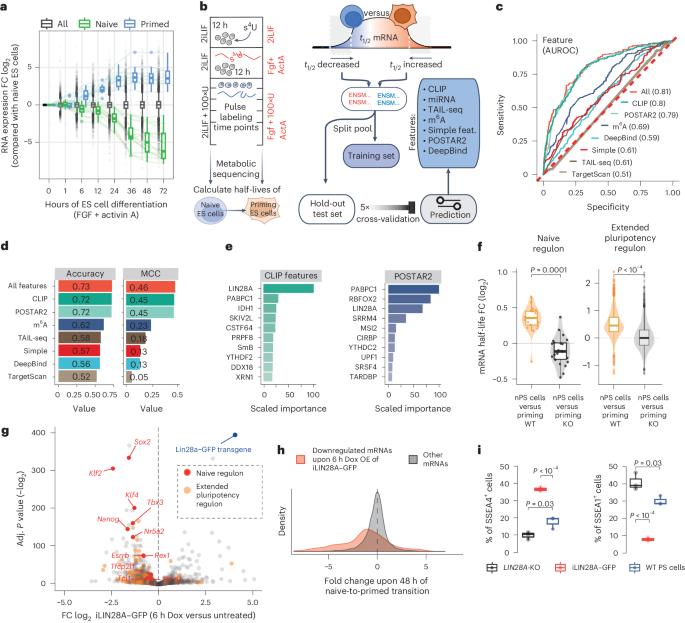
Poised PABP–RNA hubs implement signal-dependent mRNA decay in development
Signaling pathways drive cell fate transitions largely by changing gene expression. However, the mechanisms for rapid and selective transcriptome rewiring in response to signaling cues remain elusive. Here we use deep learning to deconvolve both the sequence determinants and the trans-acting regulators that trigger extracellular signal-regulated kinase (ERK)–mitogen-activated protein kinase kinase (MEK)-induced decay of the naive pluripotency mRNAs. Timing of decay is coupled to embryo implantation through ERK–MEK phosphorylation of LIN28A, which repositions pLIN28A to the highly A+U-rich 3′ untranslated region (3′UTR) termini of naive pluripotency mRNAs. Interestingly, these A+U-rich 3′UTR termini serve as poly(A)-binding protein (PABP)-binding hubs, poised for signal-induced convergence with LIN28A. The multivalency of AUU motifs determines the efficacy of pLIN28A–PABP convergence, which enhances PABP 3′UTR binding, decreases the protection of poly(A) tails and activates mRNA decay to enable progression toward primed pluripotency. Thus, the signal-induced convergence of LIN28A with PABP–RNA hubs drives the rapid selection of naive mRNAs for decay, enabling the transcriptome remodeling that ensures swift developmental progression. Here the authors show that, upon embryo implantation, signaling triggers a large-scale rearrangement of protein–RNA interactions. Phosphorylated LIN28A reassembles onto the 3′ untranslated region termini of pluripotency-associated mRNAs, where it converges with the binding of poly(A)-binding protein and drives selective mRNA decay.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Structural & Molecular Biology
BIOCHEMISTRY & MOLECULAR BIOLOGY-BIOPHYSICS
CiteScore
22.00
自引率
1.80%
发文量
160
审稿时长
3-8 weeks
期刊介绍:
Nature Structural & Molecular Biology is a comprehensive platform that combines structural and molecular research. Our journal focuses on exploring the functional and mechanistic aspects of biological processes, emphasizing how molecular components collaborate to achieve a particular function. While structural data can shed light on these insights, our publication does not require them as a prerequisite.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


