用扩展遗传字母表形成的折叠图案
IF 20.2
1区 化学
Q1 CHEMISTRY, MULTIDISCIPLINARY
引用次数: 0
摘要
在 DNA 中添加合成核苷酸可以增加 DNA 分子的线性信息密度。我们在这里报告说,它还能增加其三维折叠的多样性。具体地说,在 23 个核苷酸长的 DNA 链的 12 个位点上添加一个核苷酸(dZ,含有一个 5-硝基-6-氨基吡啶酮核碱基),就能形成一个相当稳定的单分子结构(即折叠 Z-motif,或 fZ-motif),在 pH 值为 8.5 时熔点为 66.5 °C。光谱、凝胶和二维核磁共振分析表明,折叠 Z-motif由六个反向瘦dZ-:dZ碱基对固定在一起,类似于游离杂环的晶体结构。荧光标记显示,dZ-:dZ 对以四股紧凑的下-上-下-上折叠方式连接平行链。这种结构有两种可能:一种是 dZ-:dZ 碱基对插层结构,另一种是无插层结构。夹层结构类似于 pH ≤ 6.5 时 dC:dC+ 反向配对形成的 i-motif。因此,这种 fZ 标记可能有助于 DNA 形成结合和催化所需的紧凑结构。本文章由计算机程序翻译,如有差异,请以英文原文为准。
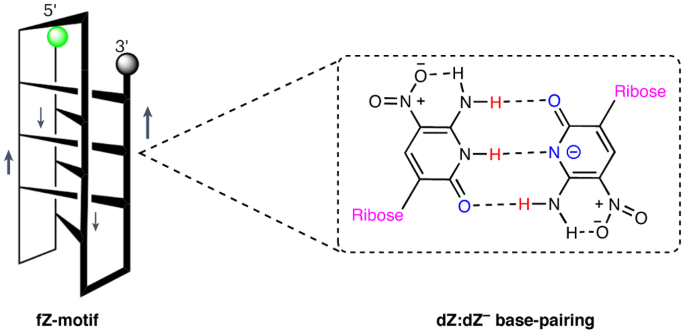

A folding motif formed with an expanded genetic alphabet
Adding synthetic nucleotides to DNA increases the linear information density of DNA molecules. Here we report that it also can increase the diversity of their three-dimensional folds. Specifically, an additional nucleotide (dZ, with a 5-nitro-6-aminopyridone nucleobase), placed at twelve sites in a 23-nucleotides-long DNA strand, creates a fairly stable unimolecular structure (that is, the folded Z-motif, or fZ-motif) that melts at 66.5 °C at pH 8.5. Spectroscopic, gel and two-dimensional NMR analyses show that the folded Z-motif is held together by six reverse skinny dZ−:dZ base pairs, analogous to the crystal structure of the free heterocycle. Fluorescence tagging shows that the dZ−:dZ pairs join parallel strands in a four-stranded compact down–up–down–up fold. These have two possible structures: one with intercalated dZ−:dZ base pairs, the second without intercalation. The intercalated structure would resemble the i-motif formed by dC:dC+-reversed pairing at pH ≤ 6.5. This fZ-motif may therefore help DNA form compact structures needed for binding and catalysis. Standard DNA is limited by low information density and functional diversity. Now it has been shown that an expanded genetic alphabet—incorporating a synthetic nucleotide, dZ—allows for the creation of stable three-dimensional DNA structures under mild alkaline conditions. Such stable structures enrich our understanding of DNA’s structural diversity and its potential in synthetic biology applications.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature chemistry
化学-化学综合
CiteScore
29.60
自引率
1.40%
发文量
226
审稿时长
1.7 months
期刊介绍:
Nature Chemistry is a monthly journal that publishes groundbreaking and significant research in all areas of chemistry. It covers traditional subjects such as analytical, inorganic, organic, and physical chemistry, as well as a wide range of other topics including catalysis, computational and theoretical chemistry, and environmental chemistry.
The journal also features interdisciplinary research at the interface of chemistry with biology, materials science, nanotechnology, and physics. Manuscripts detailing such multidisciplinary work are encouraged, as long as the central theme pertains to chemistry.
Aside from primary research, Nature Chemistry publishes review articles, news and views, research highlights from other journals, commentaries, book reviews, correspondence, and analysis of the broader chemical landscape. It also addresses crucial issues related to education, funding, policy, intellectual property, and the societal impact of chemistry.
Nature Chemistry is dedicated to ensuring the highest standards of original research through a fair and rigorous review process. It offers authors maximum visibility for their papers, access to a broad readership, exceptional copy editing and production standards, rapid publication, and independence from academic societies and other vested interests.
Overall, Nature Chemistry aims to be the authoritative voice of the global chemical community.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


