使用 xMarkerFinder 从交叉队列数据集中识别和验证微生物生物标记物。
IF 13.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
摘要
微生物特征已成为疾病诊断和预后的有前途的生物标志物,但它们在不同研究中的变异性要求生物标志物研究采用标准化方法。因此,我们引入了 xMarkerFinder,这是一个用于微生物生物标记物鉴定的四阶段计算框架,可通过跨队列数据集进行全面验证,包括差异特征鉴定、模型构建、模型验证和生物标记物解释。xMarkerFinder 可为跨队列研究鉴定和验证可重复的生物标记物,同时建立分类模型和潜在的微生物诱导机制。xMarkerFinder 最初是为肠道微生物组研究而开发的,其适应性强的设计使其适用于各种微生物栖息地和数据类型。现有的生物标记物研究工具通常只专注于一个方面,而 xMarkerFinder 则与众不同,它采用了复杂的特征选择过程,专门用于解决不同队列之间的异质性、广泛的内部和外部验证以及详细的特异性评估。执行时间取决于样本大小、所选算法和计算资源。xMarkerFinder 可通过 GitHub ( https://github.com/tjcadd2020/xMarkerFinder ) 访问,通过不同的执行选项支持不同专业水平的用户,包括带有详细教程和常见问题的分步脚本、单指令执行脚本、即用型 Docker 映像和用户友好型 Web 服务器 ( https://www.biosino.org/xmarkerfinder )。本文章由计算机程序翻译,如有差异,请以英文原文为准。
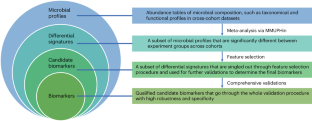
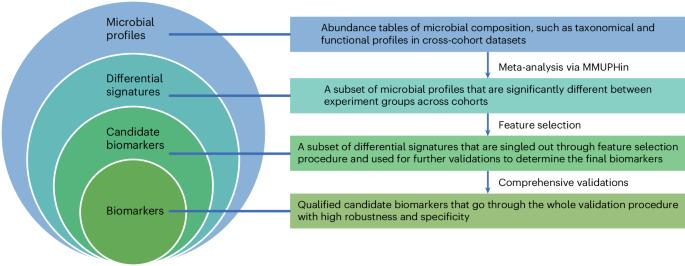
Identification and validation of microbial biomarkers from cross-cohort datasets using xMarkerFinder
Microbial signatures have emerged as promising biomarkers for disease diagnostics and prognostics, yet their variability across different studies calls for a standardized approach to biomarker research. Therefore, we introduce xMarkerFinder, a four-stage computational framework for microbial biomarker identification with comprehensive validations from cross-cohort datasets, including differential signature identification, model construction, model validation and biomarker interpretation. xMarkerFinder enables the identification and validation of reproducible biomarkers for cross-cohort studies, along with the establishment of classification models and potential microbiome-induced mechanisms. Originally developed for gut microbiome research, xMarkerFinder’s adaptable design makes it applicable to various microbial habitats and data types. Distinct from existing biomarker research tools that typically concentrate on a singular aspect, xMarkerFinder uniquely incorporates a sophisticated feature selection process, specifically designed to address the heterogeneity between different cohorts, extensive internal and external validations, and detailed specificity assessments. Execution time varies depending on the sample size, selected algorithm and computational resource. Accessible via GitHub ( https://github.com/tjcadd2020/xMarkerFinder ), xMarkerFinder supports users with diverse expertise levels through different execution options, including step-to-step scripts with detailed tutorials and frequently asked questions, a single-command execution script, a ready-to-use Docker image and a user-friendly web server ( https://www.biosino.org/xmarkerfinder ). This protocol is for using xMarkerFinder, a four-stage computational framework, to enable the identification and validation of reproducible microbial biomarkers from cross-cohort studies, and establish potential microbiome-induced mechanisms.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Nature Protocols
生物-生化研究方法
CiteScore
29.10
自引率
0.70%
发文量
128
审稿时长
4 months
期刊介绍:
Nature Protocols focuses on publishing protocols used to address significant biological and biomedical science research questions, including methods grounded in physics and chemistry with practical applications to biological problems. The journal caters to a primary audience of research scientists and, as such, exclusively publishes protocols with research applications. Protocols primarily aimed at influencing patient management and treatment decisions are not featured.
The specific techniques covered encompass a wide range, including but not limited to: Biochemistry, Cell biology, Cell culture, Chemical modification, Computational biology, Developmental biology, Epigenomics, Genetic analysis, Genetic modification, Genomics, Imaging, Immunology, Isolation, purification, and separation, Lipidomics, Metabolomics, Microbiology, Model organisms, Nanotechnology, Neuroscience, Nucleic-acid-based molecular biology, Pharmacology, Plant biology, Protein analysis, Proteomics, Spectroscopy, Structural biology, Synthetic chemistry, Tissue culture, Toxicology, and Virology.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


