端粒到端粒时代的基因组组装
IF 39.1
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
摘要
基因组序列在很大程度上决定了生物体的生物学特性并编码了生物体的历史,而从头组装--根据测序读数重建生物体基因组序列的过程--四十年来一直是生物信息学的核心问题。直到最近,基因组通常最多只能组装成几个兆字节的片段,但现在长读数测序技术的进步使许多生物的每条染色体都能进行近乎完整的组装--也称为端粒到端粒组装(telomere-to-telomere assembly)。在这里,我们回顾了最近在组装算法和协议方面取得的进展,重点是如何获得近端粒到端粒的组装。我们还讨论了解决剩余的组装差距和组装非二倍体基因组所需的其他发展。本文章由计算机程序翻译,如有差异,请以英文原文为准。
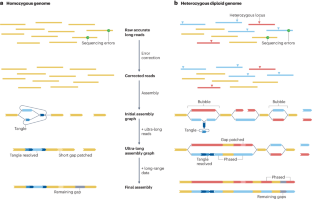
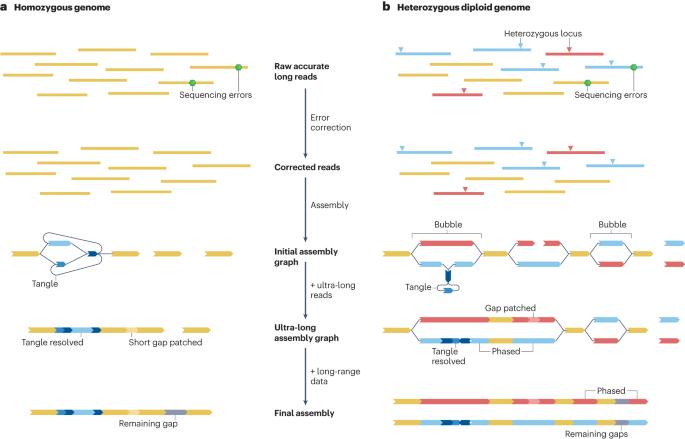
Genome assembly in the telomere-to-telomere era
Genome sequences largely determine the biology and encode the history of an organism, and de novo assembly — the process of reconstructing the genome sequence of an organism from sequencing reads — has been a central problem in bioinformatics for four decades. Until recently, genomes were typically assembled into fragments of a few megabases at best, but now technological advances in long-read sequencing enable the near-complete assembly of each chromosome — also known as telomere-to-telomere assembly — for many organisms. Here, we review recent progress on assembly algorithms and protocols, with a focus on how to derive near-telomere-to-telomere assemblies. We also discuss the additional developments that will be required to resolve remaining assembly gaps and to assemble non-diploid genomes. In this Review, Li and Durbin discuss how to generate telomere-to-telomere assemblies for large haploid or diploid genomes using currently available data types and algorithms, and outline remaining challenges in resolving highly repetitive sequences and polyploid genomes.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Nature Reviews Genetics
生物-遗传学
CiteScore
57.40
自引率
0.50%
发文量
113
审稿时长
6-12 weeks
期刊介绍:
At Nature Reviews Genetics, our goal is to be the leading source of reviews and commentaries for the scientific communities we serve. We are dedicated to publishing authoritative articles that are easily accessible to our readers. We believe in enhancing our articles with clear and understandable figures, tables, and other display items. Our aim is to provide an unparalleled service to authors, referees, and readers, and we are committed to maximizing the usefulness and impact of each article we publish.
Within our journal, we publish a range of content including Research Highlights, Comments, Reviews, and Perspectives that are relevant to geneticists and genomicists. With our broad scope, we ensure that the articles we publish reach the widest possible audience.
As part of the Nature Reviews portfolio of journals, we strive to uphold the high standards and reputation associated with this esteemed collection of publications.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


