Incongruence in the phylogenomics era
IF 39.1
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 4
Abstract
Genome-scale data and the development of novel statistical phylogenetic approaches have greatly aided the reconstruction of a broad sketch of the tree of life and resolved many of its branches. However, incongruence — the inference of conflicting evolutionary histories — remains pervasive in phylogenomic data, hampering our ability to reconstruct and interpret the tree of life. Biological factors, such as incomplete lineage sorting, horizontal gene transfer, hybridization, introgression, recombination and convergent molecular evolution, can lead to gene phylogenies that differ from the species tree. In addition, analytical factors, including stochastic, systematic and treatment errors, can drive incongruence. Here, we review these factors, discuss methodological advances to identify and handle incongruence, and highlight avenues for future research. Incongruence occurs when phylogenetic trees show conflicting evolutionary histories such as patterns of branching or relationships among taxa. This Review discusses the biological and analytical factors that lead to incongruence, methodological advances to identify and resolve incongruence, and avenues for future research.
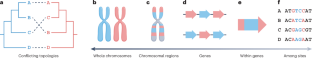
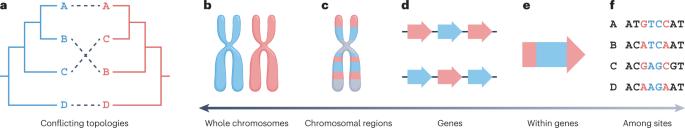
系统发育学时代的不便。
基因组规模的数据和新的统计系统发育方法的发展极大地帮助了生命树的大致草图的重建,并解决了它的许多分支。然而,不一致——对相互冲突的进化史的推断——在系统发育学数据中仍然普遍存在,阻碍了我们重建和解释生命树的能力。生物因素,如不完全谱系分类、水平基因转移、杂交、渗入、重组和趋同分子进化,可以导致不同于物种树的基因系统发育。此外,包括随机误差、系统误差和处理误差在内的分析因素也会导致不一致。在这里,我们回顾了这些因素,讨论了识别和处理不一致的方法学进展,并强调了未来研究的途径。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature Reviews Genetics
生物-遗传学
CiteScore
57.40
自引率
0.50%
发文量
113
审稿时长
6-12 weeks
期刊介绍:
At Nature Reviews Genetics, our goal is to be the leading source of reviews and commentaries for the scientific communities we serve. We are dedicated to publishing authoritative articles that are easily accessible to our readers. We believe in enhancing our articles with clear and understandable figures, tables, and other display items. Our aim is to provide an unparalleled service to authors, referees, and readers, and we are committed to maximizing the usefulness and impact of each article we publish.
Within our journal, we publish a range of content including Research Highlights, Comments, Reviews, and Perspectives that are relevant to geneticists and genomicists. With our broad scope, we ensure that the articles we publish reach the widest possible audience.
As part of the Nature Reviews portfolio of journals, we strive to uphold the high standards and reputation associated with this esteemed collection of publications.
文献相关原料
| 公司名称 | 产品信息 | 采购帮参考价格 |
|---|
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


