A chemical probe targeting the PWWP domain alters NSD2 nucleolar localization
IF 12.9
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 24
Abstract
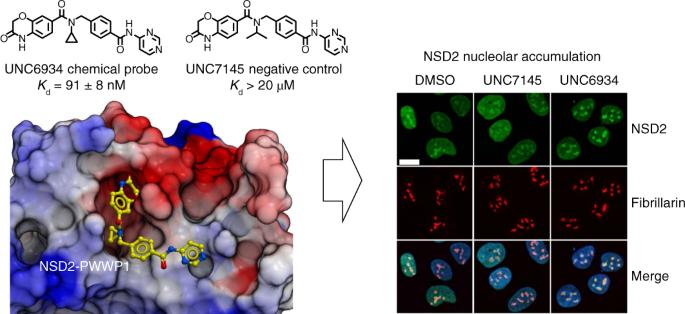
Nuclear receptor-binding SET domain-containing 2 (NSD2) is the primary enzyme responsible for the dimethylation of lysine 36 of histone 3 (H3K36), a mark associated with active gene transcription and intergenic DNA methylation. In addition to a methyltransferase domain, NSD2 harbors two proline-tryptophan-tryptophan-proline (PWWP) domains and five plant homeodomains (PHDs) believed to serve as chromatin reading modules. Here, we report a chemical probe targeting the N-terminal PWWP (PWWP1) domain of NSD2. UNC6934 occupies the canonical H3K36me2-binding pocket of PWWP1, antagonizes PWWP1 interaction with nucleosomal H3K36me2 and selectively engages endogenous NSD2 in cells. UNC6934 induces accumulation of endogenous NSD2 in the nucleolus, phenocopying the localization defects of NSD2 protein isoforms lacking PWWP1 that result from translocations prevalent in multiple myeloma (MM). Mutations of other NSD2 chromatin reader domains also increase NSD2 nucleolar localization and enhance the effect of UNC6934. This chemical probe and the accompanying negative control UNC7145 will be useful tools in defining NSD2 biology. Discovery of a chemical probe targeting the PWWP domain of NSD2 reveals insight into mechanisms that govern NSD2 localization. The compound and its negative control represent valuable tools for further defining NSD2 biology.
靶向 PWWP 结构域的化学探针可改变 NSD2 的核极定位
核受体结合含 SET 结构域的 2(NSD2)是负责组蛋白 3(H3K36)赖氨酸 36 二甲基化的主要酶,这种标记与活跃的基因转录和基因间 DNA 甲基化有关。除了一个甲基转移酶结构域外,NSD2 还含有两个脯氨酸-色氨酸-色氨酸-脯氨酸(PWWP)结构域和五个植物同源结构域(PHD),据信这些结构域是染色质阅读模块。在这里,我们报告了一种针对 NSD2 N 端 PWWP(PWWP1)结构域的化学探针。UNC6934 占据了 PWWP1 的典型 H3K36me2 结合口袋,拮抗了 PWWP1 与核糖体 H3K36me2 的相互作用,并选择性地与细胞中的内源性 NSD2 结合。UNC6934 能诱导内源性 NSD2 在核仁中聚集,从而表征多发性骨髓瘤(MM)中常见的易位导致的缺乏 PWWP1 的 NSD2 蛋白异构体的定位缺陷。其他 NSD2 染色质阅读器结构域的突变也会增加 NSD2 的核极定位,并增强 UNC6934 的作用。这种化学探针和相应的阴性对照 UNC7145 将成为确定 NSD2 生物学特性的有用工具。靶向 NSD2 PWWP 结构域的化学探针的发现揭示了 NSD2 定位的机制。该化合物及其阴性对照是进一步确定 NSD2 生物学特性的宝贵工具。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature chemical biology
生物-生化与分子生物学
CiteScore
23.90
自引率
1.40%
发文量
238
审稿时长
12 months
期刊介绍:
Nature Chemical Biology stands as an esteemed international monthly journal, offering a prominent platform for the chemical biology community to showcase top-tier original research and commentary. Operating at the crossroads of chemistry, biology, and related disciplines, chemical biology utilizes scientific ideas and approaches to comprehend and manipulate biological systems with molecular precision.
The journal embraces contributions from the growing community of chemical biologists, encompassing insights from chemists applying principles and tools to biological inquiries and biologists striving to comprehend and control molecular-level biological processes. We prioritize studies unveiling significant conceptual or practical advancements in areas where chemistry and biology intersect, emphasizing basic research, especially those reporting novel chemical or biological tools and offering profound molecular-level insights into underlying biological mechanisms.
Nature Chemical Biology also welcomes manuscripts describing applied molecular studies at the chemistry-biology interface due to the broad utility of chemical biology approaches in manipulating or engineering biological systems. Irrespective of scientific focus, we actively seek submissions that creatively blend chemistry and biology, particularly those providing substantial conceptual or methodological breakthroughs with the potential to open innovative research avenues. The journal maintains a robust and impartial review process, emphasizing thorough chemical and biological characterization.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


