Regulatory controls of duplicated gene expression during fiber development in allotetraploid cotton
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
Polyploidy complicates transcriptional regulation and increases phenotypic diversity in organisms. The dynamics of genetic regulation of gene expression between coresident subgenomes in polyploids remains to be understood. Here we document the genetic regulation of fiber development in allotetraploid cotton Gossypium hirsutum by sequencing 376 genomes and 2,215 time-series transcriptomes. We characterize 1,258 genes comprising 36 genetic modules that control staged fiber development and uncover genetic components governing their partitioned expression relative to subgenomic duplicated genes (homoeologs). Only about 30% of fiber quality-related homoeologs show phenotypically favorable allele aggregation in cultivars, highlighting the potential for subgenome additivity in fiber improvement. We envision a genome-enabled breeding strategy, with particular attention to 48 favorable alleles related to fiber phenotypes that have been subjected to purifying selection during domestication. Our work delineates the dynamics of gene regulation during fiber development and highlights the potential of subgenomic coordination underpinning phenotypes in polyploid plants. Genome and transcriptome analyses of 376 Gossypium hirsutum accessions uncover the regulation of gene expression during fiber development in allotetraploid cotton and highlight the potential for fiber quality improvement.
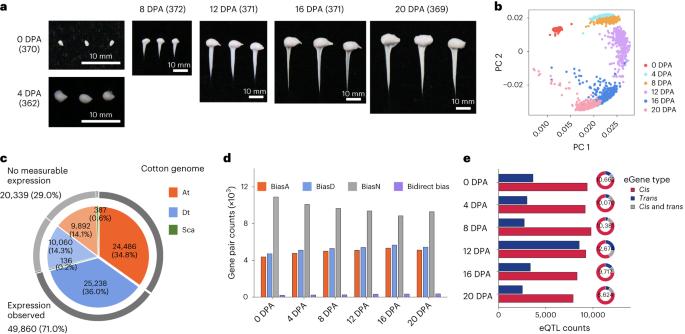
同源四倍体棉花纤维发育过程中重复基因表达的调控。
多倍体使转录调控复杂化,并增加生物体的表型多样性。多倍体中共侧子亚基因组之间基因表达的遗传调控动力学仍有待了解。本文通过对376个基因组和2215个时间序列转录组的测序,记录了异四倍体棉纤维发育的遗传调控。我们对1258个基因进行了表征,这些基因包括36个控制阶段性纤维发育的遗传模块,并揭示了控制其相对于亚基因组重复基因(同源性)的分区表达的遗传成分。只有大约30%的纤维品质相关同源性在品种中表现出表型上有利的等位基因聚集,这突出了亚基因组可加性在纤维改良中的潜力。我们设想了一种基因组育种策略,特别关注与纤维表型相关的48个有利等位基因,这些等位基因在驯化过程中经过纯化选择。我们的工作描述了纤维发育过程中基因调控的动态,并强调了亚基因组协调支持多倍体植物表型的潜力。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


