Integrated expression analysis revealed RUNX2 upregulation in lung squamous cell carcinoma tissues
IF 16.4
1区 化学
Q1 CHEMISTRY, MULTIDISCIPLINARY
引用次数: 6
Abstract
This study aimed to investigate the clinicopathological significance and prospective molecular mechanism of RUNX family transcription factor 2 (RUNX2) in lung squamous cell carcinoma (LUSC). The authors used immunohistochemistry (IHC), RNA-seq, and microarray data from multi-platforms to conduct a comprehensive analysis of the clinicopathological significance and molecular mechanism of RUNX2 in the occurrence and development of LUSC. RUNX2 expression was significantly higher in 16 LUSC tissues than in paired non-cancerous tissues detected by IHC (P < 0.05). RNA-seq data from the combination of TCGA and genotype-tissue expression (GTEx) revealed significantly higher expression of RUNX2 in 502 LUSC samples than in 476 non-cancer samples. The expression of RUNX2 protein was also significantly higher in pathologic T3-T4 than in T1-T2 samples (P = 0.031). The pooled standardised mean difference (SMD) for RUNX2 was 0.87 (95% CI, 0.58-1.16), including 29 microarrays from GEO and one from ArrayExpress. The co-expression network of RUNX2 revealed complicated connections between RUNX2 and 45 co-expressed genes, which were significantly clustered in pathways including ECM-receptor interaction, focal adhesion, protein digestion and absorption, human papillomavirus infection and PI3K-Akt signalling pathway. Overexpression of RUNX2 plays an essential role in the clinical progression of LUSC.
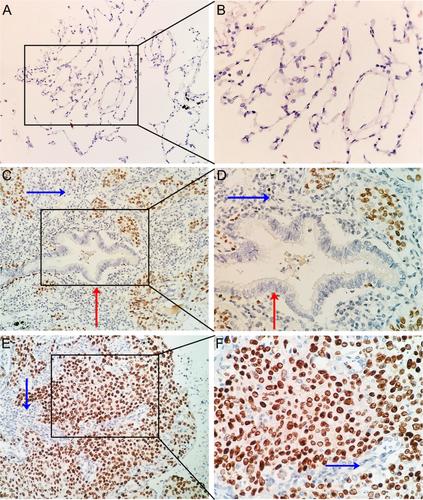
综合表达分析显示RUNX2在肺鳞癌组织中表达上调
本研究旨在探讨RUNX家族转录因子2 (RUNX2)在肺鳞癌(LUSC)中的临床病理意义及未来的分子机制。作者利用免疫组织化学(IHC)、RNA-seq、多平台微阵列数据,综合分析RUNX2在LUSC发生发展中的临床病理意义及分子机制。RUNX2在16个LUSC组织中的表达明显高于IHC检测的配对非癌组织(P < 0.05)。结合TCGA和基因型组织表达(GTEx)的RNA-seq数据显示,502例LUSC样本中RUNX2的表达明显高于476例非癌症样本。病理T3-T4组RUNX2蛋白表达也明显高于T1-T2组(P = 0.031)。RUNX2的汇总标准化平均差(SMD)为0.87 (95% CI, 0.58-1.16),包括29个来自GEO的芯片和1个来自ArrayExpress的芯片。RUNX2共表达网络揭示了RUNX2与45个共表达基因之间的复杂联系,这些基因在ecm受体相互作用、局点黏附、蛋白质消化吸收、人乳头瘤病毒感染和PI3K-Akt信号通路中显著聚集。RUNX2过表达在LUSC的临床进展中起着至关重要的作用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Accounts of Chemical Research
化学-化学综合
CiteScore
31.40
自引率
1.10%
发文量
312
审稿时长
2 months
期刊介绍:
Accounts of Chemical Research presents short, concise and critical articles offering easy-to-read overviews of basic research and applications in all areas of chemistry and biochemistry. These short reviews focus on research from the author’s own laboratory and are designed to teach the reader about a research project. In addition, Accounts of Chemical Research publishes commentaries that give an informed opinion on a current research problem. Special Issues online are devoted to a single topic of unusual activity and significance.
Accounts of Chemical Research replaces the traditional article abstract with an article "Conspectus." These entries synopsize the research affording the reader a closer look at the content and significance of an article. Through this provision of a more detailed description of the article contents, the Conspectus enhances the article's discoverability by search engines and the exposure for the research.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


