Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl
IF 50.5
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 338
Abstract
The distribution of cohesin in the mouse genome depends on CTCF, transcription and the cohesin release factor Wapl. Cohesin and CTCF are known to spatially organize mammalian genomes into chromatin loops and topologically associated domains. CTCF binds to specific DNA sequences, but it is unclear how cohesin is recruited to these sites. Here, Jan-Michael Peters and colleagues show that the distribution of cohesin in the mouse genome depends on CTCF, transcription and the cohesin-release factor Wapl. In the absence of CTCF, cohesin accumulates at the transcription start sites of active genes, which are bound by the cohesion-loading complex. In the absence of both CTCF and Wapl, cohesin accumulates at the 3′ end of active genes. The authors propose that cohesin is loaded onto DNA at sites that are distinct from its final binding sites and can be translocated by transcription until it either encounters CTCF bound to DNA or is released by Wapl. A mechanism of transcription-mediated cohesin translocation could allow the extrusion of chromatin loops. Mammalian genomes are spatially organized by CCCTC-binding factor (CTCF) and cohesin into chromatin loops1,2 and topologically associated domains3,4,5,6, which have important roles in gene regulation1,2,4,5,7 and recombination7,8,9. By binding to specific sequences10, CTCF defines contact points for cohesin-mediated long-range chromosomal cis-interactions1,2,4,5,6,7,11. Cohesin is also present at these sites12,13, but has been proposed to be loaded onto DNA elsewhere14,15 and to extrude chromatin loops until it encounters CTCF bound to DNA16,17,18,19. How cohesin is recruited to CTCF sites, according to this or other models, is unknown. Here we show that the distribution of cohesin in the mouse genome depends on transcription, CTCF and the cohesin release factor Wings apart-like (Wapl). In CTCF-depleted fibroblasts, cohesin cannot be properly recruited to CTCF sites but instead accumulates at transcription start sites of active genes, where the cohesin-loading complex is located14,15. In the absence of both CTCF and Wapl, cohesin accumulates in up to 70 kilobase-long regions at 3′-ends of active genes, in particular if these converge on each other. Changing gene expression modulates the position of these ‘cohesin islands’. These findings indicate that transcription can relocate mammalian cohesin over long distances on DNA, as previously reported for yeast cohesin20,21,22,23, that this translocation contributes to positioning cohesin at CTCF sites, and that active genes can be freed from cohesin either by transcription-mediated translocation or by Wapl-mediated release.
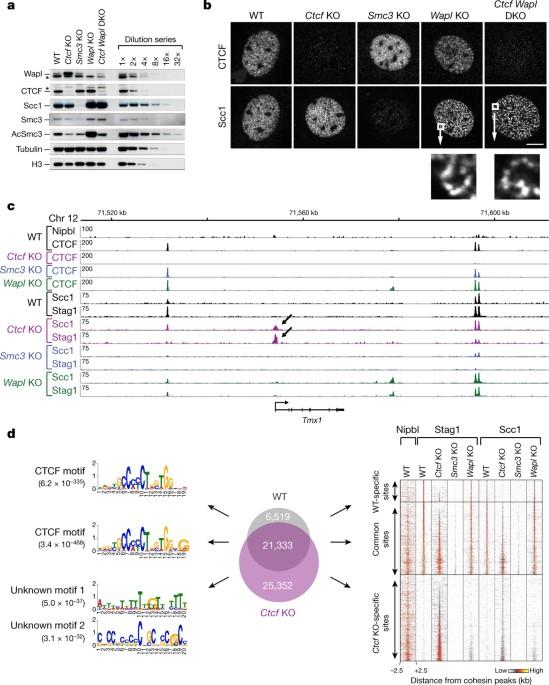
聚合素通过转录、CTCF 和 Wapl 在哺乳动物基因组中定位。
哺乳动物基因组由 CCCTC 结合因子(CTCF)和粘合素在空间上组织成染色质环和拓扑相关域,它们在基因调控和重组中发挥着重要作用。通过与特定序列结合,CTCF 为凝聚素介导的长程染色体顺式相互作用定义了接触点。粘合素也存在于这些位点,但有人认为粘合素会被加载到其他地方的 DNA 上,并挤出染色质环,直到遇到与 DNA 结合的 CTCF。根据这一模型或其他模型,凝聚素如何被招募到 CTCF 位点尚不清楚。在这里,我们发现,小鼠基因组中凝聚素的分布取决于转录、CTCF 和凝聚素释放因子 Wings apart-like (Wapl)。在缺失 CTCF 的成纤维细胞中,粘合素不能被正常招募到 CTCF 位点,而是聚集在活跃基因的转录起始位点,而粘合素负载复合物就位于该位点。在缺乏 CTCF 和 Wapl 的情况下,凝聚素会聚集在活性基因 3'- 端长达 70 千碱基的区域,尤其是当这些区域相互交汇时。基因表达的改变会调节这些 "凝聚素岛 "的位置。这些研究结果表明,转录可以将哺乳动物的凝聚素在DNA上进行长距离迁移,就像以前报道的酵母凝聚素一样,这种迁移有助于将凝聚素定位在CTCF位点上,而且活性基因可以通过转录介导的迁移或Wapl介导的释放从凝聚素中释放出来。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature
综合性期刊-综合性期刊
CiteScore
90.00
自引率
1.20%
发文量
3652
审稿时长
3 months
期刊介绍:
Nature is a prestigious international journal that publishes peer-reviewed research in various scientific and technological fields. The selection of articles is based on criteria such as originality, importance, interdisciplinary relevance, timeliness, accessibility, elegance, and surprising conclusions. In addition to showcasing significant scientific advances, Nature delivers rapid, authoritative, insightful news, and interpretation of current and upcoming trends impacting science, scientists, and the broader public. The journal serves a dual purpose: firstly, to promptly share noteworthy scientific advances and foster discussions among scientists, and secondly, to ensure the swift dissemination of scientific results globally, emphasizing their significance for knowledge, culture, and daily life.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


