DNA-binding-domain fusions enhance the targeting range and precision of Cas9
IF 36.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 113
Abstract
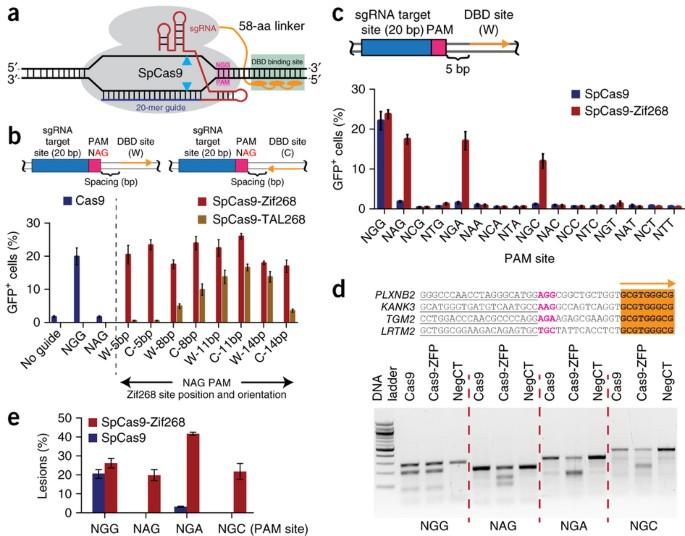
Fusing a DNA-binding domain to Cas9 with an attenuated, more promiscuous PAM-recognition domain increases the targeting range of Cas9 as well as its specificity. The CRISPR-Cas9 system is commonly used in biomedical research; however, the precision of Cas9 is suboptimal for applications that involve editing a large population of cells (for example, gene therapy). Variations on the standard Cas9 system have yielded improvements in the precision of targeted DNA cleavage, but they often restrict the range of targetable sequences. It remains unclear whether these variants can limit lesions to a single site in the human genome over a large cohort of treated cells. Here we show that by fusing a programmable DNA-binding domain (pDBD) to Cas9 and attenuating Cas9’s inherent DNA-binding affinity, we were able to produce a Cas9-pDBD chimera with dramatically improved precision and an increased targeting range. Because the specificity and affinity of this framework can be easily tuned, Cas9-pDBDs provide a flexible system that can be tailored to achieve extremely precise genome editing at nearly any genomic locus.
DNA结合域融合增强了Cas9的靶向范围和精度
将 DNA 结合结构域与减弱的、更杂乱的 PAM 识别结构域融合到 Cas9 中,增加了 Cas9 的靶向范围及其特异性。CRISPR-Cas9 系统常用于生物医学研究;然而,对于涉及编辑大量细胞的应用(如基因治疗)来说,Cas9 的精确性并不理想。标准 Cas9 系统的变体提高了靶向 DNA 切割的精确度,但它们往往限制了靶向序列的范围。目前仍不清楚这些变体是否能将病变限制在人类基因组中的单个位点上,从而影响大量细胞。在这里,我们展示了通过将可编程DNA结合域(pDBD)与Cas9融合并减弱Cas9固有的DNA结合亲和力,我们能够制备出一种Cas9-pDBD嵌合体,其精确性得到了显著提高,靶向范围也扩大了。由于这种框架的特异性和亲和力很容易调整,Cas9-pDBDs 提供了一种灵活的系统,几乎可以在任何基因组位点实现极其精确的基因组编辑。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Methods
生物-生化研究方法
CiteScore
58.70
自引率
1.70%
发文量
326
审稿时长
1 months
期刊介绍:
Nature Methods is a monthly journal that focuses on publishing innovative methods and substantial enhancements to fundamental life sciences research techniques. Geared towards a diverse, interdisciplinary readership of researchers in academia and industry engaged in laboratory work, the journal offers new tools for research and emphasizes the immediate practical significance of the featured work. It publishes primary research papers and reviews recent technical and methodological advancements, with a particular interest in primary methods papers relevant to the biological and biomedical sciences. This includes methods rooted in chemistry with practical applications for studying biological problems.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


