POKY software tools encapsulating assignment strategies for solution and solid-state protein NMR data
IF 5.1
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 5
Abstract
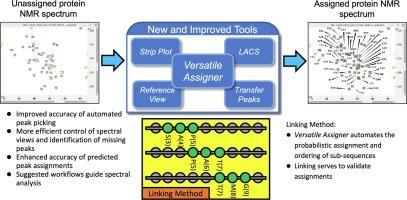
NMR spectroscopy provides structural and functional information about biomolecules and their complexes. The complexity of these systems can make the NMR data difficult to interpret, particularly for newer users of NMR technology, who may have limited understanding of the tools available and how they are used. To alleviate this problem, we have created software based on standardized workflows for both solution and solid-state NMR spectroscopy of proteins. These tools assist with manual and automated peak picking and with chemical shift assignment and validation. They provide users with an optimized path through spectral analysis that can help them perform the necessary tasks more efficiently.
POKY软件工具封装分配策略的溶液和固态蛋白质核磁共振数据
核磁共振波谱提供了生物分子及其复合物的结构和功能信息。这些系统的复杂性可能使NMR数据难以解释,特别是对于NMR技术的新用户来说,他们可能对可用的工具及其使用方式了解有限。为了缓解这个问题,我们创建了基于标准化工作流程的软件,用于蛋白质的溶液和固态核磁共振波谱。这些工具有助于手动和自动峰采摘,以及化学班次分配和验证。它们通过光谱分析为用户提供优化的路径,可以帮助他们更有效地执行必要的任务。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of Structural Biology: X
Biochemistry, Genetics and Molecular Biology-Structural Biology
CiteScore
6.50
自引率
0.00%
发文量
20
审稿时长
62 days
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


