Single-cell DNA methylome and 3D genome atlas of human subcutaneous adipose tissue
IF 29
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
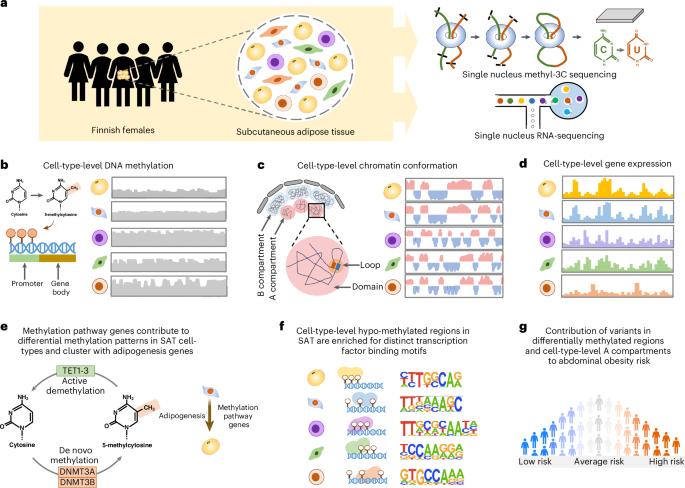
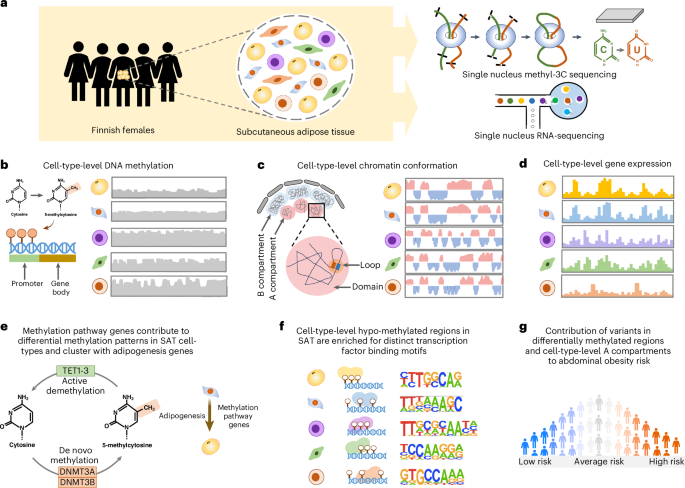
The cell-type-level epigenomic landscape of human subcutaneous adipose tissue (SAT) is not well characterized. Here, we elucidate the epigenomic landscape across SAT cell types using snm3C-seq. We find that SAT CG methylation (mCG) displays pronounced hypermethylation in myeloid cells and hypomethylation in adipocytes and adipose stem and progenitor cells, driving nearly half of the 705,063 differentially methylated regions (DMRs). Moreover, TET1 and DNMT3A are identified as plausible regulators of the cell-type-level mCG profiles. Both global mCG profiles and chromosomal compartmentalization reflect SAT cell-type lineage. Notably, adipocytes display more short-range chromosomal interactions, forming complex local 3D genomic structures that regulate transcriptional functions, including adipogenesis. Furthermore, adipocyte DMRs and A compartments are enriched for abdominal obesity genome-wide association study (GWAS) variants and polygenic risk, while myeloid A compartments are enriched for inflammation. Together, we characterize the SAT single-cell-level epigenomic landscape and link GWAS variants and partitioned polygenic risk of abdominal obesity and inflammation to the SAT epigenome. This multiomic study, including single-nucleus DNA methylation and chromatin conformation matched with single-nuclei RNA sequencing, provides insights into the epigenomic landscape of human subcutaneous adipose tissue.
人皮下脂肪组织单细胞DNA甲基组和三维基因组图谱
人类皮下脂肪组织(SAT)的细胞类型水平的表观基因组景观尚未得到很好的表征。在这里,我们使用snm3C-seq阐明了SAT细胞类型的表观基因组景观。我们发现,SAT CG甲基化(mCG)在髓细胞中表现出明显的高甲基化,在脂肪细胞、脂肪干细胞和祖细胞中表现出低甲基化,驱动了705,063个差异甲基化区域(DMRs)的近一半。此外,TET1和DNMT3A被认为是细胞型水平mCG谱的合理调节因子。全球mCG谱和染色体区隔化都反映了SAT细胞类型谱系。值得注意的是,脂肪细胞表现出更多的短程染色体相互作用,形成复杂的局部3D基因组结构,调节转录功能,包括脂肪生成。此外,脂肪细胞DMRs和A区室在腹部肥胖全基因组关联研究(GWAS)变异和多基因风险中富集,而髓细胞A区室在炎症中富集。总之,我们描述了SAT单细胞水平的表观基因组景观,并将GWAS变异和腹部肥胖和炎症的分割多基因风险与SAT表观基因组联系起来。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


