Integrative genomic reconstruction reveals heterogeneity in carbohydrate utilization across human gut bifidobacteria
IF 19.4
1区 生物学
Q1 MICROBIOLOGY
引用次数: 0
Abstract
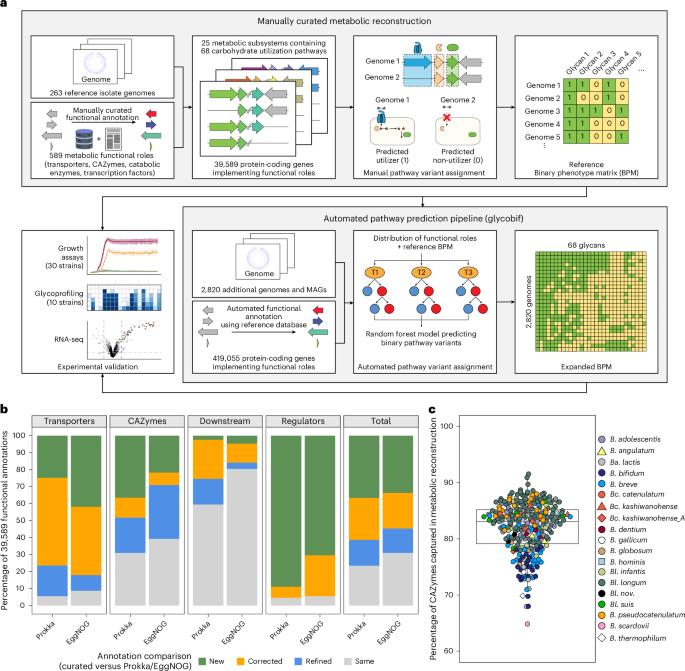
Bifidobacteria are beneficial saccharolytic microbes that are widely used as probiotics or in synbiotic formulations, yet individual responses to supplementation can vary with strain type, microbiota composition, diet and lifestyle, underscoring the need for strain-level insights into glycan metabolism. Here we reconstructed 68 pathways for the utilization of mono-, di-, oligo- and polysaccharides by analysing the distribution of 589 curated metabolic gene functions (catabolic enzymes, transporters and transcriptional regulators) across 3,083 non-redundant Bifidobacterium genomes of human origin. Thirty-eight predicted phenotypes were validated in vitro for 30 geographically diverse strains, supporting genomics-based predictions. Our analysis uncovered extensive inter- and intraspecies functional heterogeneity, including a distinct clade within Bifidobacterium longum that metabolizes α-glucans and Bangladeshi isolates carrying unique gene clusters for xyloglucan and human milk oligosaccharide utilization. This large-scale genomic compendium advances our understanding of bifidobacterial carbohydrate metabolism and can inform the rational design of probiotic and synbiotic formulations tailored to strain-specific nutrient preferences. A comprehensive genomic analysis reveals species- and strain-level heterogeneity in carbohydrate utilization potential across bifidobacteria of human origin.

整合基因组重建揭示了人类肠道双歧杆菌碳水化合物利用的异质性
双歧杆菌是有益的溶糖微生物,被广泛用作益生菌或合成制剂,但个体对补充剂的反应可能因菌株类型、微生物群组成、饮食和生活方式而异,这强调了菌株水平对聚糖代谢的了解的必要性。本研究通过分析3083个非冗余双歧杆菌人类基因组中589个调控代谢基因功能(分解代谢酶、转运蛋白和转录调节因子)的分布,重建了68个利用单、二、寡和多糖的途径。38种预测表型在30个地理位置不同的菌株中进行了体外验证,支持基于基因组学的预测。我们的分析揭示了广泛的种间和种内功能异质性,包括长双歧杆菌中代谢α-葡聚糖的独特分支,以及孟加拉国分离的携带木葡聚糖和人乳低聚糖利用的独特基因簇。这种大规模的基因组纲要促进了我们对双歧杆菌碳水化合物代谢的理解,并可以为根据菌株特定营养偏好量身定制的益生菌和合成制剂的合理设计提供信息。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Microbiology
Immunology and Microbiology-Microbiology
CiteScore
44.40
自引率
1.10%
发文量
226
期刊介绍:
Nature Microbiology aims to cover a comprehensive range of topics related to microorganisms. This includes:
Evolution: The journal is interested in exploring the evolutionary aspects of microorganisms. This may include research on their genetic diversity, adaptation, and speciation over time.
Physiology and cell biology: Nature Microbiology seeks to understand the functions and characteristics of microorganisms at the cellular and physiological levels. This may involve studying their metabolism, growth patterns, and cellular processes.
Interactions: The journal focuses on the interactions microorganisms have with each other, as well as their interactions with hosts or the environment. This encompasses investigations into microbial communities, symbiotic relationships, and microbial responses to different environments.
Societal significance: Nature Microbiology recognizes the societal impact of microorganisms and welcomes studies that explore their practical applications. This may include research on microbial diseases, biotechnology, or environmental remediation.
In summary, Nature Microbiology is interested in research related to the evolution, physiology and cell biology of microorganisms, their interactions, and their societal relevance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


