Design of PROTACs utilizing the E3 ligase GID4 for targeted protein degradation
IF 10.1
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
Proteolysis targeting chimeras (PROTACs) hijack E3 ligases and the ubiquitin–proteasome system to achieve selective degradation of neo-substrates. Their ability to target otherwise intractable substrates has rendered them a valuable modality in drug discovery. However, only a handful of over 600 human E3 ligases have been functionalized for PROTAC applications. Here we show that the E3 ligase GID4 (glucose-induced degradation deficient complex 4) can be leveraged for targeted protein degradation using a noncovalent small molecule. We design and synthesize GID4-based PROTACs, exemplified by NEP162, which can eliminate endogenous BRD4 in a GID4- and ubiquitin–proteasome system-dependent manner. NEP162 exhibits antiproliferative activity and inhibits tumor growth in a xenograft model, hinting toward potential anticancer applications. We further present the crystal structures of GID4–PROTAC–BRD4 ternary complexes in three distinct states, unveiling plastic interactions between GID4 and BRD4. These structural insights, combined with in vitro and in vivo data, decipher the molecular basis by which the hereby developed PROTACs recruit BRD4 to GID4 for targeted degradation and expand our arsenal of PROTAC-exploitable E3 ligases. Here the authors show that the E3 ligase GID4 can be harnessed for targeted protein degradation and present the crystal structure of the GID4–PROTAC–BRD4 ternary complex to elucidate the underlying molecular mechanisms.
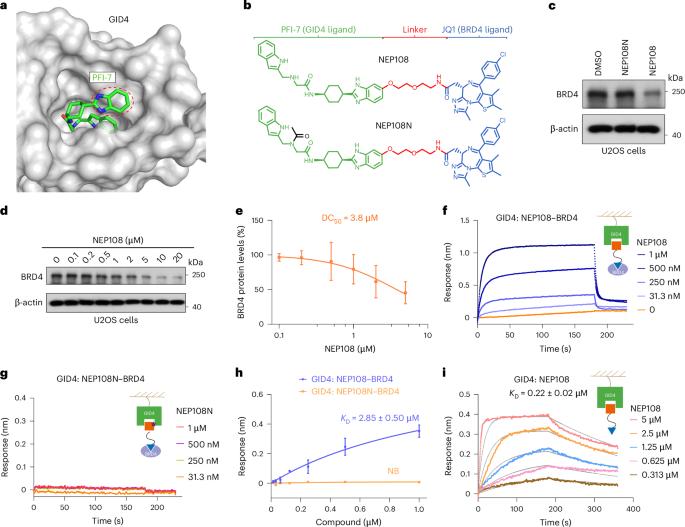

利用E3连接酶GID4进行靶向蛋白降解的PROTACs设计
蛋白水解靶向嵌合体(PROTACs)劫持E3连接酶和泛素-蛋白酶体系统来实现新底物的选择性降解。它们靶向其他难以处理的底物的能力使它们成为药物发现的一种有价值的方式。然而,在600多个人类E3连接酶中,只有少数被功能化用于PROTAC应用。在这里,我们展示了E3连接酶GID4(葡萄糖诱导的降解缺陷复合物4)可以利用非共价小分子进行靶向蛋白质降解。我们设计并合成了基于GID4的PROTACs,以NEP162为例,它可以以GID4和泛素蛋白酶体系统依赖的方式消除内源性BRD4。在异种移植物模型中,NEP162显示出抗增殖活性并抑制肿瘤生长,这暗示了其潜在的抗癌应用。我们进一步展示了GID4 - protac - BRD4三元配合物在三种不同状态下的晶体结构,揭示了GID4和BRD4之间的塑性相互作用。这些结构见解,结合体外和体内数据,破译了由此开发的PROTACs招募BRD4到GID4进行靶向降解的分子基础,并扩展了我们的protac可利用的E3连接酶库。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Structural & Molecular Biology
BIOCHEMISTRY & MOLECULAR BIOLOGY-BIOPHYSICS
CiteScore
22.00
自引率
1.80%
发文量
160
审稿时长
3-8 weeks
期刊介绍:
Nature Structural & Molecular Biology is a comprehensive platform that combines structural and molecular research. Our journal focuses on exploring the functional and mechanistic aspects of biological processes, emphasizing how molecular components collaborate to achieve a particular function. While structural data can shed light on these insights, our publication does not require them as a prerequisite.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


